GeWorkbench Release 2.1
From Informatics
General notes on previous geWorkbench releases
- General notes, feature requests and FAQ page - This page was started with material from the time of release 1.7.0 and will be updated continually.
- Links to other release pages:
- The geWorkbench Roadmap (local version) contains possible directions for future development.
- caBIG has a separate geWorkbench Roadmap page that we must maintain.
- caBIG/NCI also provides the official download page for geWorkbench
Release Schedule for 2.1.0
- geWorkbench 2.1.0 code freeze: August 13th, 2010 (planned)
- System Testing concluded: August 20th, 2010
- Final release target: September 3rd, 2010
- Actual release date: September 10th, 2010
Role Assignments
- Release Manager – Kenneth Smith
- Release Engineer – Thomas Garben
- Tech Lead – Zhou Ji
- Tester – Udo Többen, and the rest of the bunch
- Test Manager – Udo Többen
- Technical Writer – Mary VanGinhoven
Things to remember
- Best practices for defect management - See also Aris's email of 8/20/09 on this topic.
- geWorkbench Roadmap page at NCICB - keep up to date with actual plans and developments - at https://cabig-kc.nci.nih.gov/Molecular/KC/index.php/GeWorkbench_Roadmap
- InstallAnywhere JRE update packs: http://www.flexerasoftware.com/products/installanywhere/files-utilities.htm
Known Issues - General
- InstallAnywhere and Norton Internet Security Sonar - Under Windows, InstallAnywere places a file called "Install.exe" in a folder in a path like "C:\Users\ksmith\AppData\Local\Temp\I1276186086\Windows\. This file was seen to be detected and removed by "Norton Sonar", silently terminating the install. Everything below "Temp" is removed after the installation finishes.
Known Issues - release 2.1
See the 2.0 release page for existing issues. Need to see how many of those got fixed.
New Component Detail and Dependencies pages created
Annotation Dependencies - list of dependencies of particular components on particular annotation file columns.
CNKB Data - release status and available interactions for each database.
Major changes in release 2.1.0
Major Code Changes in 2.1.0
A number of components had major code cleanups:
List of changes to GUI
New components in release 2.1
- Coefficient of Variation filter.
Other major new features in release 2.1
- bug 2323 and others - Enhancements to BLAST (in sequence alignment).
- bug 2340 - New Feature - System information display.
List of other major changes
As of 8/11/2010, 37 bugs are marked resolved, closed, or closed pending documentation. 48 issues are still open, of which some will be resolved for release 2.1.0.
- bug 2303 - Fixed GO tree display in CNKB component. The tree is now expandable.
- bug 2242 - In filtering components, check for valid inputs.
- bug 2190 - Fixed problem when saving a workspace that contains a large dendrogram display from hierarchical clustering.
- bug 2305 - Fixed a problem with JMOL performance.
- bug 2294 - Welcome screen is now version aware - it will always appear first time when a new version is run.
Documentation
Documentation changes pending from previous releases
See the 2.0 release page for a long list of pending documentation changes and other TODO items.
Tutorial/Online Help chapters that need coordinated updating before release
- Arrays component - http://wiki.c2b2.columbia.edu/workbench/index.php/Data_Subsets_-_Arrays#Lower_Pane
- Save - not implemented in release 2.0 or earlier. Implemented in 2.1 development version.
- Markers component - says Save is not implemented but I don't think this is right.
Tutorial/Online Help coordinated changes to both finished
- ARACNe - updated discussion of upper limit on arrays, it pertains only to Fixed Bandwidth. Also made same changes in Online Help.
- MINDy - added description of how to use ARACNe preprocessing. Full resynch of Tutorial and Online Help.
- Filtering - added Coefficient of Variation.
- Pattern Discovery - Existing tutorial (revised during release 1.8.0) ported to Online Help for release 2.1.0.
Versions of external files/components included in this release
- gene_ontology.1_2.obo downloaded 8/10/2010 from geneontology.org.
- Ontologizer.jar version 2.0, file created 3/10/2010 (as seen by inspecting files in the JAR file). We are using the "Command line" jar file. No change from release 2.0.0. Checked no further updates as of 8/10/2010. http://compbio.charite.de/index.php/ontologizer2.html
- Note - On 5/31/2010, the Ontologizer "Manual" version jar file (which has a GUI) was updated. However, the command line version was still not updated.
- Jaspar_CORE (http://jaspar.genereg.net/) - Unchanged from release 2.0.0. SQL files last updated on server 10/2009. (/html/DOWNLOAD/jaspar_CORE/non_redundant/all_species/sql_tables)
- JMOL - JMOL 12 RC.10. No change from release 2.0.0. Jar file last updated 5/13/2010.
System Testing Notes
How to test BLAST queries
Our BLAST component uses the URLAPI - that is, it sends the query to NCBI BLAST as a string of arguments embedded in a URL. We can then test our Blast GUI by inspecting the resulting URL string. To see this string, one must change a log setting. In the geWorkbench installation directory, find conf/log4j.properties. Within this file, towards the bottom, add this line under "#component packages:"
- log4j.logger.org.geworkbench.components.alignment=DEBUG
Components
List of Included Components
Noted on SVN refresh on 8/11/2010 - a number of outdated components look like dropped from SVN. See 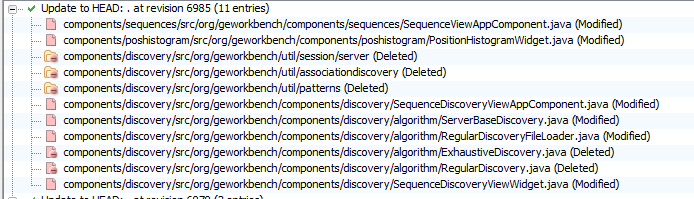
Data Managmenent:
- Arrays/Phenotypes
- Markers
- Preferences
- Project Panel
- Session manager - no one knows what this is - probably a SOAP interface. But it is definitely needed!
File input formats
- Affy File Format
- CEL File Loader
- Exp. Format
- FASTA Format
- Genepix File Format
- PDB Structure Format
- Tab-delimited (RMA Express Format)
Connectivity
- caArray2 - updated to support caArray 2.3.0 in release 1.8.0 (released September 2009). The caArray client jar is NOT backwards-compatible with any previous versions.
Data filters
- Filtering
- Affy Detection Call Filter
- Coefficient of Variation (new)
- Deviation Filter
- Expression Threshold Filter
- Genepix Filter (Two channel filter)
- Genepix Flag Filter
- Missing Values Filter
Normalization
- HouseKeeping Genes Normalizer
- Normalization
- Log2 Tranformation
- Marker Centering Normalizer
- Mean Variance Normalizer
- Missing Values (Normalizer)
- Microarray Centering Normalizer
- Quantile Normalizer
- Threshold Normalizer
Experiment Information
- Dataset Annotation
- Dataset History
- Experiment Info
- Version Information
Analyis/Visualization
- Alignment Results
- Analysis
- ANOVA
- ARACNe2 - adds Adaptive Partitioning algorithm and Preprocessing mode.
- caBIO Pathways (this has been integrated in the Marker Annotations component)
- Cancer Gene Index integration in the Marker Annotations component.
- CELImageViewer
- Cellular Networks Knowledge Base
- Color Mosaic
- Component Configuration Manager.
- Cytoscape_V2_4 - updated version of Cytoscape.
- Dendrogram
- Expression Profiles
- Expression Value Distribution
- Gene Ontology Enrichment Analysis and Display
- Hierarchical Clustering Analysis
- genSpace collaborative framework
- Image Viewer
- Jmol
- Marker Annotations
- MarkUs - Analysis and Viewer
- MRA - Master Regulator Analysis
- MatrixREDUCE
- Microarray Viewer
- MINDy - Analysis and Viewer
- Pattern Discovery
- Position Histogram
- Pudge?? - Analysis and Viewer (Browser) - if this is working (Kiran?) we should include. We can create a very simple online help file, essentially pointing to the Pudge documentation at the Honig site (Aris).
- Promoter
- Scatter Plot
- Sequence
- Sequence Alignment
- Sequence Retriever
- SOM Analysis
- SOM Clusters
- t Test Analysis
- Tabular Microarray Viewer
- Volcano Plot
- GenePattern components
- PCA (GenePattern) - Analysis and Viewer
- K-nearest neighbors (GenePattern)
- SVM 3.0 (GenePattern) - Analysis and Viewer - include, we need to develop online help and tutorial (Aris).
- WV - Weighted Voting (GenePattern)
- GeneWays (need to update status for release 2.0) (need for Cytoscape viewer for ARACNe)- this component not working; we do
not have end-user documentation materials (Aris). However, Geneways must be included for Cytoscape interaction to function, but Geneways itself cannot be chosen as a component nor used directly.
Excluded and Dropped Components
The release creation script in build.xml now explicitly includes components by name (previously it excluded components by name) The following is a list of modules known to be excluded.
Excluded components
The following components are excluded for a variety of reasons, most often due to lack of formal requirements documentation or/and associated system test scripts. Some of them should be scheduled for inclusion in the next production release. For modules not found in the current all.xml a path to the component is shown.
Still under development:
- CART (GenePattern) - this component has not yet been released. Is part of another component and must be excluded manually from the final installer release build.
- Cancer-GEMS (awaiting further development from NCI)
- NetBoost
- EdgeListFileFormat (NetBoost)
- Evidence Integration
- MEDUSA
Not actively being developed:
- GCRMA Via R CEL Loader (in \geworkbench\src\org\geworkbench\components\parsers)
- GSEA
- Multi-t-test (OK, but need to understand when it would be used, e.g. after ANOVA, and if it is what we really want).
- SMLR - Sparse Multinomial Logistic Regression - implementation by John Watkinson.
- SVM Format (in \geworkbench\src\org\geworkbench\components\parsers) (left over from a John Watkinson project).
- Synteny (in \geworkbench\components\alignment\src\org\geworkbench\components\alignment\client)
- t-profiler
- caScript
Dropped components
These components are not expected to be used again.
- CuteNet (GeneWays)
- Column Major Format (in \geworkbench\src\org\geworkbench\components\parsers)
- Frequency Threshold Filter (There is a class called AllelicFrequencyThresholdFilter in \geworkbench\components\filtering\src\org\geworkbench\components\filtering)
- GeneOntology (the original component, now replaced by geneontology2/Ontologizer2.0)
- Genotypic File Format (in \geworkbench\src\org\geworkbench\components\parsers\genotype)
- Network Browser (was part of Reverse Engineering - would require major rewrite to revive. PathwayDecoder is module name)
- Pattern Discovery Algorithm (association analysis)
- Patterns (Pattern Panel) - Omit from release - Appears to have been superseded by the Sequence component.
- Reverse Engineering (non-ARACNE, unpublished algorithm. PathwayDecoder is module name)
- Simulation (a student project)
Note - the original "interactions" component was dropped and reimplemented as the Cellular Networks Knowledge Base. It took a brief detour as being called component "interactions2".
Externally supplied components
The following components originate external to the geWorkbench source tree:
MatrixReduce
Source
MatrixReduce source code was obtained from the Bussemaker lab and a modified copy saved under: adcvs.cu-genome.org:/cvs/magnet/matrixreduce_distribution. This modified copy contains Java API changes made to integrate with geWorkbench.
Compiling
MatrixReduce is compiled using the following commands:
- FitModel binary is compiled manually as follows
- gcc -c -O2 -mno-cygwin -funroll-loops *.c
- gcc -mno-cygwin -static nrutil.o fncs_cmns.o fncs_seqs.o fncs_tdat.o fncs_seed.o fncs_app1.o fncs_app2.o fncs_nrcs.o fncs_topo.o fncs_mylm.o fncs_bits.o FitModel.o -o FitModel –lm (for windows and linux)
- gcc -mno-cygwin nrutil.o fncs_cmns.o fncs_seqs.o fncs_tdat.o fncs_seed.o fncs_app1.o fncs_app2.o fncs_nrcs.o fncs_topo.o fncs_mylm.o fncs_bits.o FitModel.o -o FitModelMac –lm (for Mac)
- API jar: The Java API jar is created with the makefile, command "make jar".
- FitModel binary is compiled manually with gcc, with extra flags to tell it to not use Cygwin, to optimize and to unroll loops
- FitModel.exe bundles both the NR (Numerical Recipies) and GNU libraries.
- The API jar is created with the makefile under MatrixREDUCE's top directory.
Notes
See comment on white spaces in file names/paths in Mantis : http://mantis.cu-genome.org/view.php?id=1316
Aracne.jar for MINDY
Although ARACNE is a geWorkbench component, the MINDY component uses a version of ARACNE that is externally maintained. The file aracne.jar is copied directly into the geWorkbench CVS tree.
The location of the external ARACNE code is:
The version of the external ARACNE code is:
Cytoscape
Any other components?
Analysis components - external runtime dependencies
- has not been updated for 2.1.
| component | local | external type | username/password | relay servlet | known to work outside campus |
|---|---|---|---|---|---|
| ANOVA | yes | grid | grid_default | no | ? |
| ARACNe | yes | grid | grid_default | no | ? |
| CNKB | no | servlet | some open data | yes | ? |
| MINDy | yes | grid | grid_default | no | ? |
| GenSpace | local | grid | genSpace account | no | ? |
| Hierarchical Clustering | yes | grid | grid_default | no | ? |
| KNN | no | GenePattern | ??? | no | ? |
| MarkUs | no | grid | open | no | ? |
| MRA | local | no | - | no | not applicable |
| MatrixREDUCE | local | grid | grid_default | no | ? |
| PCA | no | GenePattern | ??? | no | ? |
| PUDGE | no | web | open | no | ? |
| SkyLine | no | grid | grid_default | no | ? |
| SkyBase | no | grid | grid_default | no | ? |
| SOM | yes | grid | grid_default | no | ? |
| SVM | no | GenePattern | ??? | no | ? |
| WV | no | GenePattern | ??? | no | ? |
Important links
http://www.psl.cs.columbia.edu/genspace/