GeWorkbench-web/ANOVA
|
Home | Overview | Set View | File Formats | Desktop Tutorials |
ANOVA | ARACNe | Cellular Networks KnowledgeBase | Gene Ontology | Hierarchical Clustering | MarkUs | msViper | T-Test |
Contents
Overview
The ANOVA (ANalysis of VAriance) algorithm (Zar, 1999) is used to determine whether any significant difference in the means exist in a dataset composed of three or more groups of experimental tests.
The geWorkbench ANOVA component implements a one-way analysis of variance calculation derived from TIGR's MeV (MultiExperiment Viewer) (Saeed, 2003). At least three groups of arrays must be specified by defining and activating them in the ANOVA "Array Sets" selector. For each marker the routine determines if, at the specified level of significance, any difference in the mean exists in expression values between any of the array sets (the null hypothesis is that there is no difference between the groups). Several basic methods of multiple testing correction are offered. The analysis does not indicate between which groups the difference is found, only that one exists.
Those markers for which a significant difference is found are placed into a new set in the main Marker Sets list (under "Set View") called "Significant Genes". The results are also displayed in a tabular viewer.
Setting up an ANOVA run
Prerequisites
- A microarray expression dataset must already be loaded into the Workspace.
- At least three array sets must already be defined in the Array Sets/Phenotypes list under "Set View".
Selecting Case and Control Arrays
- Marker Context - If more than one context for marker sets has been created, first choose the desired context. Each context can contain its own collection of marker sets.
- Limit to selected markers (optional) - Selecting marker sets here allows the analysis to be limited to the markers in the union of the selected sets.
- Array Context - If more than one context for array sets has been created, first choose the desired context. Each context can contain its own collection of array sets.
- Select array sets for comparison - Choose at least three array sets to be compared by ANOVA. No array should belong to more than one of the chosen sets.
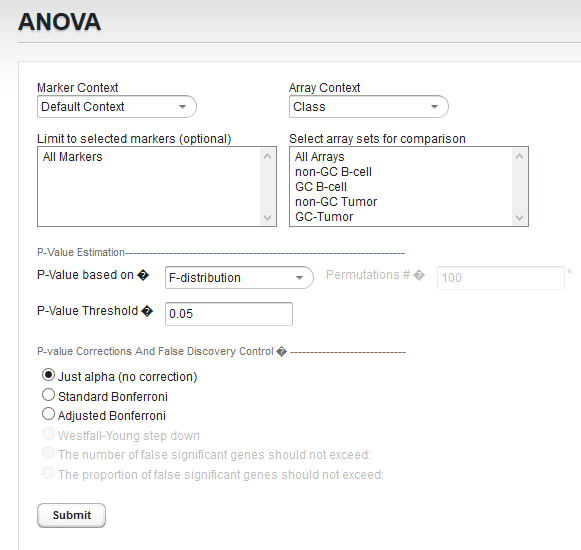
ANOVA Parameters and Settings
P-Value Estimation
The P-value represents, for any one test (one marker), the probability of falsely rejecting the null hypothesis - that is, calling a difference real when it is not. It is the probability that an F-statistic a least as large as obtained would be occur under the null hypothesis of no difference in means.
- P-value based on - Select one of two methods for calculating p-values:
- F-Distribution - The p-value will be calculated using the F-distribution. The F-distribution arises from the ratio of the variances of two normally distributed statistics (chi-squared distributions).
- Permutation - Permutations of the data will be used to generate a distribution against which the significance of the observed difference is judged. The number of desired permutations can be entered. The default number of permutations is 100.
- P-value threshold - sets the value of alpha, the critical p-value, for judging whether the null hypothesis can be rejected - that is, whether a difference is regarded as significant.
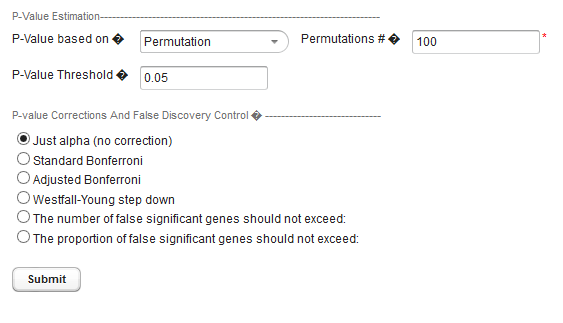
The below image shows that three additional P-value corrections become enabled when the permutation method is chosen.
P-value corrections
Several methods for correcting for the effects of performing multiple tests are offered, including Bonferroni and False Discovery Rate control. They differ in how they compare the calculated p-value to the cutoff value of alpha - the critical p-value for determining the significance of an observed difference.
- Just Alpha - No correction is performed.
- Standard Bonferroni - The cutoff value (alpha) is divided by the number of tests (genes) before being compared with the calculated p-values.
- Adjusted (step-down) Bonferroni - Similar to the Bonferroni correction, but for each successive P-value in a list of p-values sorted in increasing order, the divisor for alpha is decremented by one and then the result compared with the P-value. The effect is to slightly reduce the stringency (increase the power) of the Bonferroni correction. This is a step-down procedure.
- Westfall-Young Step-Down - (Dudoit, 2003) Another step-down procedure which adjusts the critical value alpha using the Max T method. (This correction is only available when the permutation method is chosen for calculating p-values).
False Discovery Control
(This correction is only available when the permutation method is chosen for calculating p-values).
Rather than controlling the family-wise error rate (FWER) as do the Bonferroni corrections, that is, the probability of even one false positive occurring in the multiple trials, the false discovery rate calculation controls the rate of false positives. This can result in increased power to detect true differences. See Korn, 2001 and Korn, 2004, if one can accept more false positives. The number of false positives that is acceptable may be an economic decision, based on how many follow-up tests can be performed.
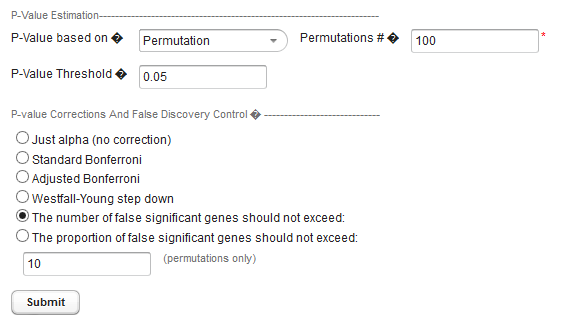
The user must select a limit to the rate of false discoveries as follows and enter the cutoff value in the text field below the method section radio buttons (shown in figure above):
- The number of false significant genes should not exceed - An upper limit on the number of false positives (markers falsely called as showing a significant difference), or
- The proportion of false significant genes should not exceed - An upper limit on the proportion of false positives.
Analysis Actions
- Submit - start the ANOVA analysis
Example of running ANOVA
Loading Data
This example uses the microarray data file Bcell-100_log2.exp, which is the Bcell data described in the Tutorial_Data section, and which has been further normalized as follows:
- Threshold Normalization - set a minimum value of 1.0 for each data point,followed by
- Log2 transformation
Briefly, this dataset is composed of 100 Affymetrix HG-U95Av2 arrays on which various B-cell lines, both normal and cancerous, were analyzed. Thus it explores a potentially wide variety of expression phenotypes.
- Load the Bcell-100_log2.exp dataset into geWorkbench as type "Expression File (.exp)".
- Load/associate the Affymetrix HG-U95Av2 annotation file if desired. A copy is preloaded into geWorkbench-web and can be associated with the data file during the upload process.
Setting Parameters
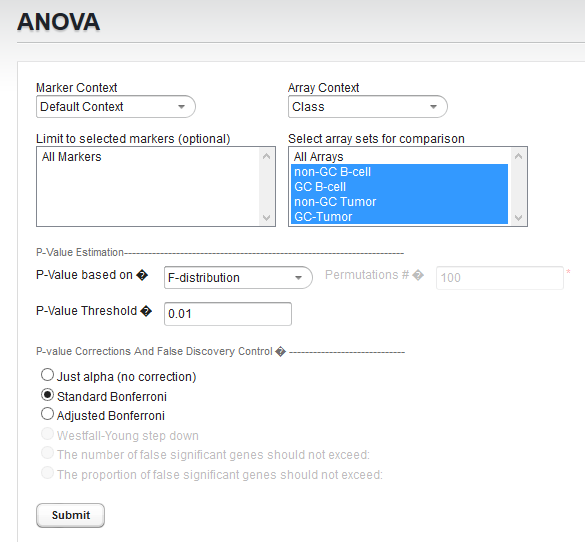
Choosing array groups
The Bcell-100_log2.exp dataset comes with predefined sets of arrays.
- In the pulldown under "Array Contex", select the context "Class".
- In the list under "Select array sets for comparison" , choose all four available array sets.
Set Parameters and submit ANOVA job
For this example we will apply a relatively stringent multiple testing correction.
- Leave the P-value method set to F-distribution.
- Leave the P-Value Threshold (alpha) set to 0.01.
- For the P-value correction choose Standard Bonferroni.
- Push the Submit button.
ANOVA Results
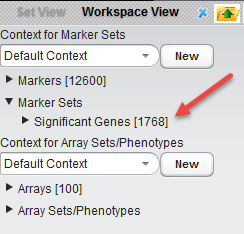
Significant markers set
All markers which met the threshold p-value (alpha) cutoff are placed into a "Significant Genes" set under Marker Sets in the Set View. Such sets of markers can be used as the starting point for further characterization and analysis.
The ANOVA result node in the Workspace
When the ANOVA calculation completes, an "Anova" result node is placed in the Workspace. When the result node is selected (highlighted), the results will be displayed in tabular form.
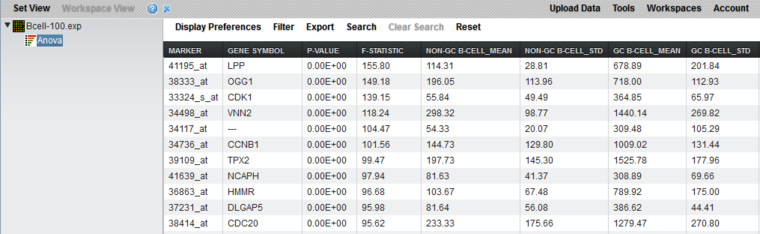
Tabular Viewer
This Visual Area component displays a read-only spreadsheet view of the significant genes sorted by p-value in ascending order (from most significant to least significant).
Spreadsheet columns
- Marker Name - Shows the gene name if an annotation file has been loaded, otherwise shows the probeset name.
- P-value - the probability of observing an F-statistic this large by chance alone, assuming the null hypothesis of no actual differences between sets of arrays. If a multiple testing correction (e.g. Bonferroni) was used, the corrected p-value is reported.
- F-statistic - the raw ANOVA score for each marker.
- Mean - the mean expression value for each group of arrays.
- Std - the standard deviation for each group of arrays.
Control Menus
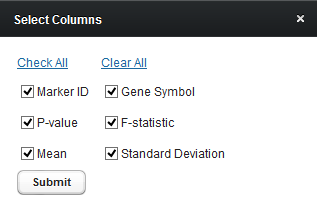
Display Preferences
Controls which of the columns to display. The choices, described in the previous section, are Marker ID, Gene Symbol, P-value, F-statistic, Mean, and Standard Deviation.
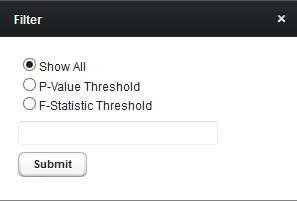
Filter
Limit the tabular display based on the following choices
- Show All
- P-Value Threshold
- F-statistic Threshold
Export
Export the results table to a CSV format file. The exported file will contain only the columns displayed.
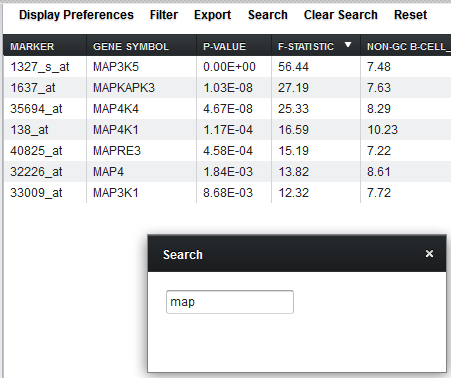
Search
Dynamically search for a marker name or gene symbol. The display updates as characters are entered in the search box. The table has been sorted on F-statistic by clicking on the column header.
Clear Search
Clear the search result and return to the normal tabular display.
Reset
Reset will undo any changes made to the display, such as sorting, searching, or filtering.
Further customizing the tabular view
- Resize columns by using the mouse to drag column boundaries.
- Sort the spreadsheet on a specific column by double clicking on its header. Succesive clicks will toggle between ascending order and descending order.
Data Information
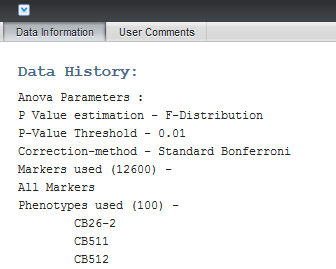
Details about each run are maintained in the Dataset History. With the ANOVA result node highlighted in the Workspace, the Dataset History display can be seen by clicking on the small arrow at lower left in the display. It includes the following information:
- P-Value estimation method
- P-Value threshold
- Multiple testing correction method
- Complete list of arrays (phenotypes) in each group analyzed
- Complete list of all markers analyzed.
References
TIGR MeV lists the following relevant citations
- Dudoit S., J.P. Shaffer and J.C. Boldrick 2003. Multiple Hypothesis Testing in Microarray Experiments. Statistical Science 18: 71-103
- Korn, E.L., J.F. Troendle, L.M. McShane, R. Simon (2001).Controlling the number of false discoveries: application to high-dimensional genomic data. Technical report 003, Biometric Research Branch, National Cancer Institute. http://linus.nci.nih.gov/~brb/TechReport.htm
- Korn, E.L., J.F. Troendle, L.M. McShane, R. Simon (2004).Controlling the number of false discoveries: application to high-dimensional genomic data. Journal of Statistical Planning and Inference 124: 379-398.
- Saeed AI, Sharov V, White J, Li J, Liang W, Bhagabati N, Braisted J, Klapa M, Currier T, Thiagarajan M, Sturn A, Snuffin M, Rezantsev A, Popov D, Ryltsov A, Kostukovich E, Borisovsky I, Liu Z, Vinsavich A, Trush V, Quackenbush J. TM4: a free, open-source system for microarray data management and analysis. Biotechniques. 2003 Feb;34(2):374-8. PMID 12613259
- Zar, J.H. 1999. Biostatistical Analysis. 4th ed. Prentice Hall, NJ., pp 178-182.