CeRNA Query
|
Home | Quick Start | Basics | Menu Bar | Preferences | Component Configuration Manager | Workspace | Information Panel | Local Data Files | File Formats | caArray | Array Sets | Marker Sets | Microarray Dataset Viewers | Filtering | Normalization | Tutorial Data | geWorkbench-web Tutorials |
Analysis Framework | ANOVA | ARACNe | BLAST | Cellular Networks KnowledgeBase | CeRNA/Hermes Query | Classification (KNN, WV) | Color Mosaic | Consensus Clustering | Cytoscape | Cupid | DeMAND | Expression Value Distribution | Fold-Change | Gene Ontology Term Analysis | Gene Ontology Viewer | GenomeSpace | genSpace | Grid Services | GSEA | Hierarchical Clustering | IDEA | Jmol | K-Means Clustering | LINCS Query | Marker Annotations | MarkUs | Master Regulator Analysis | (MRA-FET Method) | (MRA-MARINa Method) | MatrixREDUCE | MINDy | Pattern Discovery | PCA | Promoter Analysis | Pudge | SAM | Sequence Retriever | SkyBase | SkyLine | SOM | SVM | T-Test | Viper Analysis | Volcano Plot |
Contents
Overview
This component provides query access to a precomputed database of competitive endogenous RNA (ceRNA) interactions, also called "sponge" interactions. These interactions underlie a post-transcriptional layer of regulation, and were predicted using the Hermes algorithm (Sumazin et al., 2011) with subsequent improvements (submitted).
Datasets for Glioblastoma (GBM), Ovarian (OV), Breast (BRCA), and Prostate (PRAD) cancer are currently available. Each is calculated by analyzing gene expression profiles in combination with matched microRNA (miR) profiles obtained from the TCGA.
Hermes predicts the intersection set (program) of miRNAs which bind to both of two different RNAs. Variation in the level of one of the RNA species may affect the level of the second RNA via its effects on the availability of the various miRNAs.
Controls
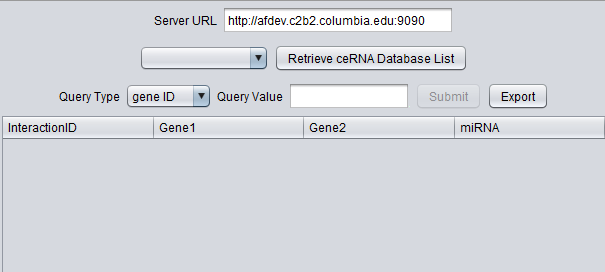
The figure below shows the basic query interface.
- Server URL - The URL for the ceRNA server should be entered.
- Retrieve ceRNA Database List - push to populate the pulldown menu of database choices.
- Query Type - Gene ID or miRNA ID. Select whether to search on a Gene ID (gene symbol) or a miRNA ID.
- Query Value - Enter the desired search term.
- Submit - perform the query.
- Export - Export the query result to a CSV-format file.
Output
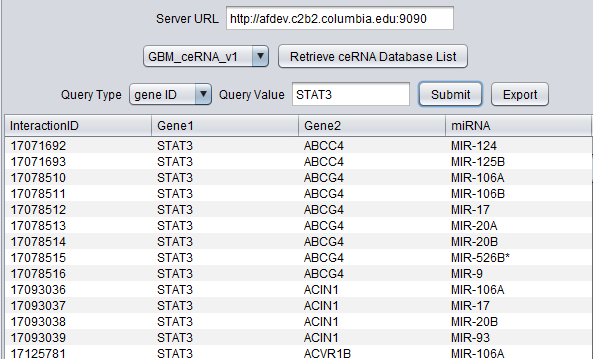
Querying on a gene symbol, STAT3:
The data columns returned are:
- InteractionID - internal record number from the CNKB database.
- Gene1 and Gene2 - the two competing RNA molecules.
- miRNA - a miRNA which is predicted to bind to both RNA molecules.
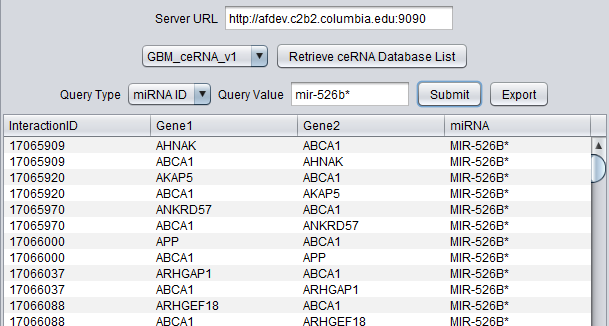
Querying on a miRNA ID (Note - * indicates the less-used strand):
References
- Sumazin P, Yang X, Chiu HS, Chung WJ, Iyer A, Llobet-Navas D, Rajbhandari P, Bansal M, Guarnieri P, Silva J, Califano A. (2011) An extensive microRNA-mediated network of RNA-RNA interactions regulates established oncogenic pathways in glioblastoma. Cell 147(2):370-81. doi: 10.1016/j.cell.2011.09.041 (link to paper).
- Hermes website for application download.