Color Mosaic
Contents
Overview
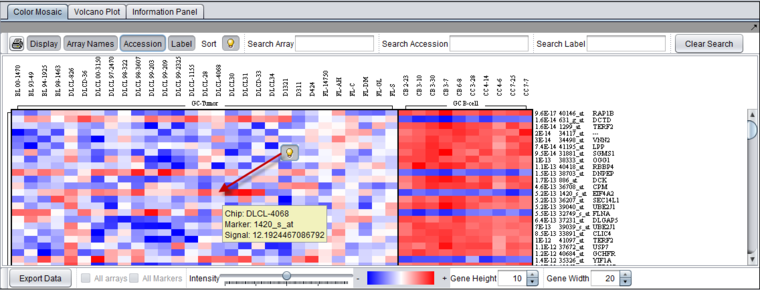
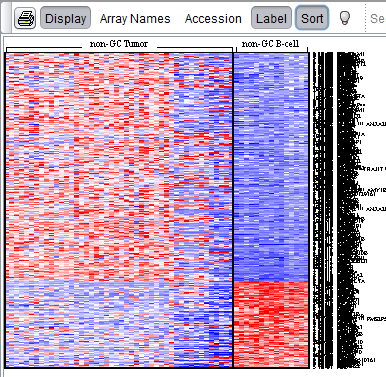
The Color Mosaic component displays expression values for all active arrays and markers using an "Eisen" plot "heat map" colored by either an absolute or relative color scheme. It is located in the View Area ( see figure below) in the top-right of the application.
Measurements in the selected microarray dataset are displayed as a two-dimensional matrix with each column representing an array and each row representing a gene/marker. The color spectrum is used to indicate the magnitudes of the measurements.
When the Color Mosaic component is used to display the results of a t-test or ANOVA, it will display the p-values to the right of each marker.
Limitations
There is a limit of about 100 Megapixels to the size of the Color Mosaic image that can be displayed. A warning will be issued if this size is exceeded, and the image will not be created.
Color Mosaic Controls
The figure above shows the Color Mosaic component displaying the results of a t-test, including p-values.
Display Controls
- Display - Toggles on/off the Color Mosaic display. It is better to leave the display "off" when not being used.
- Array Names - Toggles on/off the display of array names above the columns of the color mosaic.
- Accession - Toggles on/off the display of the accession number (e.g. Affymetrix probeset name) to the right of each row in the color mosaic.
- Label - Toggles on/off the display of the gene symbol(if available) to the right of each row in the color mosaic.
- Sort - The "Sort" control is disabled unless the results of an analysis are being displayed. It toggles between two sorting orders for the t-test results. These are further described below. It is not used by other analyses.
- Lightbulb
 - Activates the tooltip function over the color mosaic display. When activated, mouse-over (hover) text over any cell in the color mosaic will give the following information:
- Activates the tooltip function over the color mosaic display. When activated, mouse-over (hover) text over any cell in the color mosaic will give the following information:
- Chip - the name of the array
- Marker - the marker name/accession number (e.g. Affymetrix probeset id).
- Signal - The value of the signal in the dataset for the marker/array pair.
- Intensity - The intensity slider increases or decreases the thresholds used to define the color mappings, as explained in the Color Key section.
- Gene Height and Gene Width - Change the size (number of pixels) assigned to each marker (row height) or array (column width) in the color mosaic. The associated labels scale proportionately.
Search Controls
There are separate text boxes to search for array names, accession numbers, and labels. The search fields are enabled by their respective buttons.
Each search field features progressive search. As each character is typed in, the first match to the entered string will be automatically highlighted in blue. The next match can be found by hitting the "Enter" key. Typing additional characters further narrows the search.
- Search Array - Toggle on/off searching for array names.
- Search Accession - Toggle on/off searching for marker IDs.
- Search Label - Toggle on/off searching for Gene Names.
- Clear Search - Clear the search result highlights.
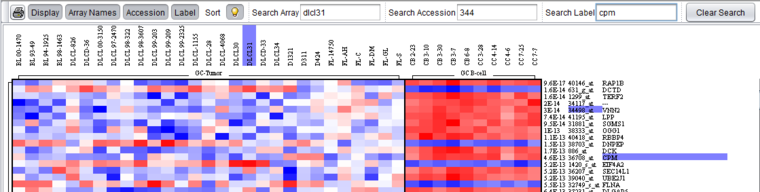
The figure below shows the results, with labels highlighted in purple, of searches using all three search fields.
Sort button
The Sort button toggles between two methods of sorting the markers in the Color Mosaic component. It is enabled only when displaying results from a t-test.
- By default, sorting is by p-value, with the smallest values at top.
- If the Sort button is pushed, the following alternate sorting method is used:
- Divide the markers into two groups based on the sign of the t-value.
- Sort each group by t-value in ascending order.
- Recombine the two groups such that the group with increasingly positive t-values comes first, followed by the second group with decreasingly negative t-values.
This leaves the most significant positive and and negative t-values adjacent in the center of the sorted list"
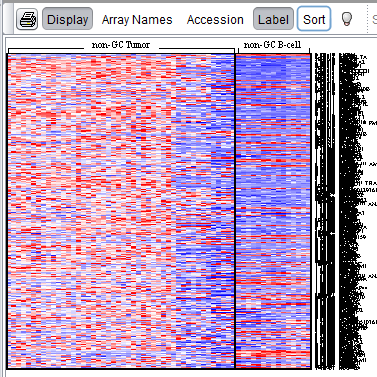
A t-test result displayed default sorted by p-value:
and the same result displayed sorted by the alternate method (Sort button activated):
Other controls
- Export data - This option is only available when displaying results of an analysis, e.g. from ANOVA or t-test. Export p-values to a file.
- All Markers - Overrides any set of markers activated in the Markers component to display all markers. Disabled in some displays such as ANOVA.
- All Arrays - Overrides any set of arrays activated in the Arrays/Phenotypes component to display results from all arrays. Disabled in some displays such as ANOVA.
- Printer Icon - Send the heat-map to the printer.
Mouse-click actions
Left-click
Left-clicking on any location (tile) in the mosaic will highlight the corresponding gene probe in the Markers component, and the corresponding array in the Arrays/Phenotypes component.
Right-click
Right-clicking anywhere on the color mosaic display will bring up a menu with the following entries, as shown in the below figure:
- Print - Send the heat map to the printer.
- Zoom-in/ Zoom-out - Increase/decrease size of color mosaic.
- Take Snapshot - Save an image of the heat map to the Workspace.
- Export significance value - Export p-values from displayed ANOVA / t-test result. (Disabled when not displaying an analysis result).
Color key
The Color key shows the range of colors available in the currently set Visualization preference (Menu->Tools->Preferences->Visualization). There are two available color schemes: Absolute and Relative. In each, the lowest and highest expression values correspond to the most saturated colors, and values closer to the middle are indicated by shading into a third color.
See Visualization preference for switching between absolute and relative color schemes.
Absolute
Colors are calculated on an absolute scale centered about zero (when the Intensity Slider is in the middle, neutral position). Values shade between the following:
- Black - zero.
- Red - positive values.
- Green - negative values.
Relative
Colors are calculated relative to the mean for each marker (when the Intensity Slider is in the middle, neutral position). Expression value colors are directly comparable only within a particular marker.
- White - mean value for marker.
- Red - values greater than the mean.
- Blue - values less than the mean.