Difference between revisions of "SAM"
(→Local Installation of R Server) |
|||
| Line 15: | Line 15: | ||
Setting the R location in geWorkbench is covered at [[Preferences#R_Location_.28Rscript.exe_and_its_folder.29| Preferences: R Location]]. | Setting the R location in geWorkbench is covered at [[Preferences#R_Location_.28Rscript.exe_and_its_folder.29| Preferences: R Location]]. | ||
| + | |||
| + | |||
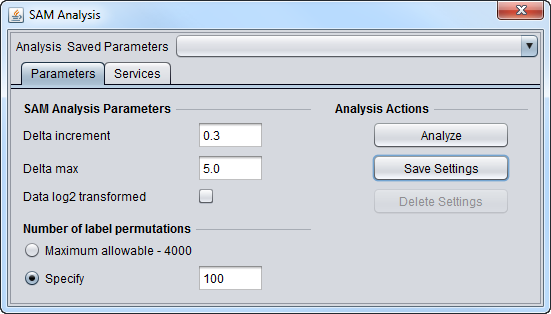
| + | ===SAM Parameters=== | ||
| + | |||
| + | [[Image:SAM_Analysis_Parameters.png]] | ||
| + | |||
| + | |||
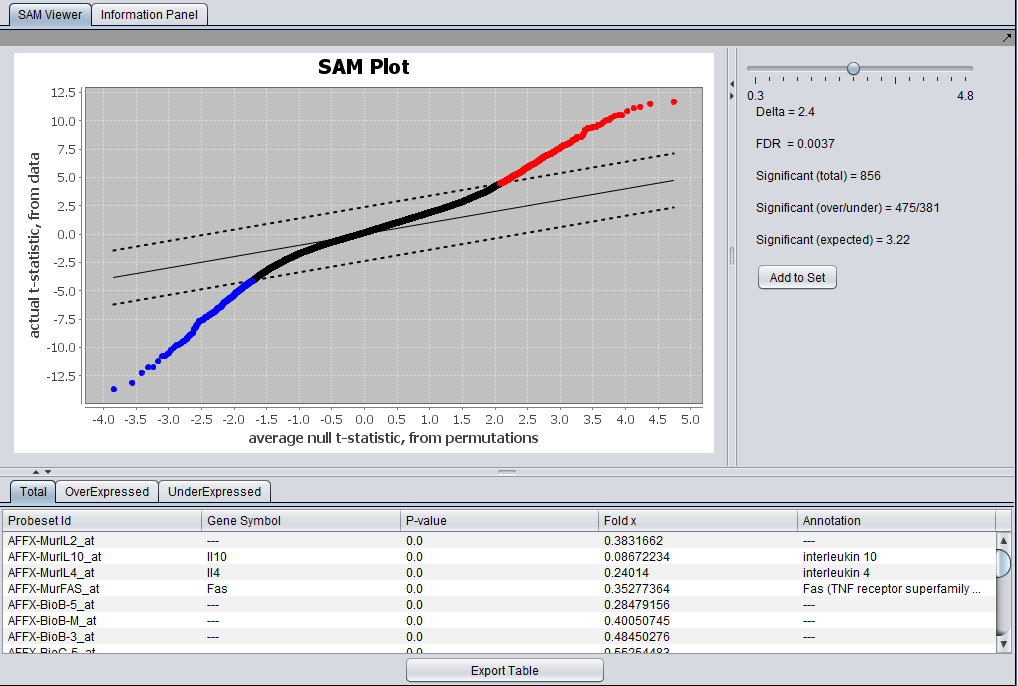
| + | ===SAM Results Viewer=== | ||
| + | The Full SAM Reults viewer is shown here. | ||
| + | |||
| + | [[Image:SAM_Viewer_full.png]] | ||
| + | |||
| + | |||
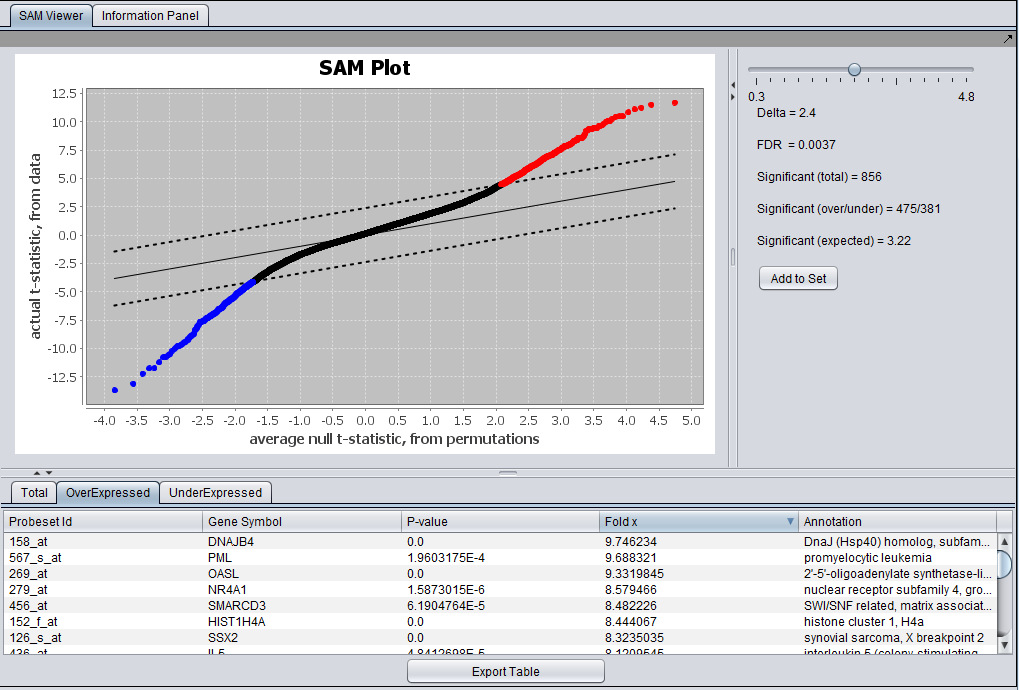
| + | The results can be sorted, e.g. on the fold change value as shown below. | ||
| + | |||
| + | [[Image:SAM_Viewer_Overexpressedl.png]] | ||
Revision as of 17:48, 22 October 2013
Contents
SAM - Significance Analysis of Microarrays
Local Installation of R Server
The SAM analysis component can use a local installation of R on your desktop computer.
This has been tested with R version 2.15.0 and 3.0.2.
There are special considerations for installing R on Windows computers, please see R installation on Windows.
Setting the R location in geWorkbench is covered at Preferences: R Location.
SAM Parameters
SAM Results Viewer
The Full SAM Reults viewer is shown here.
The results can be sorted, e.g. on the fold change value as shown below.