Difference between revisions of "T-test"
| Line 7: | Line 7: | ||
In this tutorial, you will: | In this tutorial, you will: | ||
| − | * Get acquainted with the | + | * Get acquainted with the t-Test and Multi t-Test |
| − | * Apply a | + | * Apply a t-Test and Multi t-Test |
| Line 16: | Line 16: | ||
| − | == | + | ==t-Test== |
| − | + | A t-Test analysis can be used to identify markers with statistically significant differential expression between sets of microarrays. The t-test determines, for each marker, if there is a significant difference between the two groups (case and control). To perform this analysis, you must classify the sets, set the analysis parameters and view the results in the visualization components. A detailed description of the t-Test parameters is described in online help. | |
| Line 25: | Line 25: | ||
This process has already been described in [[Tutorial - Data Subsets]]. Briefly, | This process has already been described in [[Tutorial - Data Subsets]]. Briefly, | ||
| − | 1. Mark the '''Cardio''' phenotype a 'Case'. By default, | + | 1. Mark the '''Cardio''' phenotype a 'Case'. By default, sets are marked as control. Sets classified Case are shown with a red thumbtack icon. |
* Right-click on '''Cardio''' phenotype. | * Right-click on '''Cardio''' phenotype. | ||
* Select '''Classification'''>'''Case'''. | * Select '''Classification'''>'''Case'''. | ||
| − | 2. Activate the arrays '''Normal''' and '''CCMP''' by selecting the checkboxes next to the | + | 2. Activate the arrays '''Normal''' and '''CCMP''' by selecting the checkboxes next to the set name. |
| Line 46: | Line 46: | ||
| − | === | + | ===t-Test Results=== |
| Line 54: | Line 54: | ||
|- | |- | ||
|- | |- | ||
| − | | Markers which met the significance test are included in a new | + | | Markers which met the significance test are included in a new Marker Set called “Significant Genes”. || [[Image:E_ttestgpanel.png]] |
|- | |- | ||
|- | |- | ||
| Line 62: | Line 62: | ||
| − | The values of the | + | The values of the t-Test can be seen in the Color Mosaic panel and the Volcano Plot. |
| Line 75: | Line 75: | ||
|- | |- | ||
|- | |- | ||
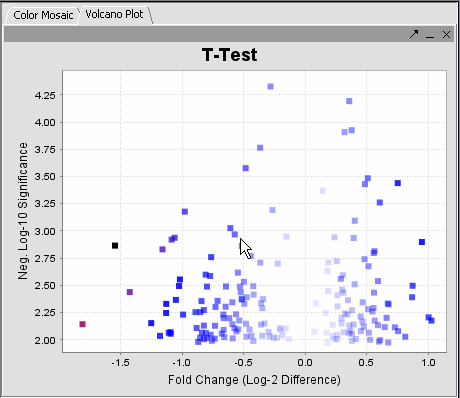
| − | | Clicking on any of the spots highlights the marker selected in the Marker | + | | Clicking on any of the spots highlights the marker selected in the Marker component. * Insert another description || |
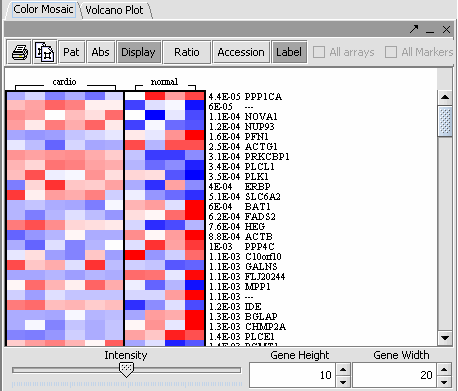
* The label to the right displays the Significance value ( lower the value, most likely different) and gene name for the displayed genes. The genes are displayed in ascending order by Significance Value. | * The label to the right displays the Significance value ( lower the value, most likely different) and gene name for the displayed genes. The genes are displayed in ascending order by Significance Value. | ||
| Line 88: | Line 88: | ||
* Display: Must be toggled on to display data. | * Display: Must be toggled on to display data. | ||
| − | * ''Pat, Abs, Ratio Overlapping Pages | + | * ''Pat, Abs, Ratio and Overlapping Pages Icons: These are not relevant to the t-Test display.'' |
|- | |- | ||
|- | |- | ||
|} | |} | ||
| + | |||
| + | ==Multi t-test== | ||
| + | |||
| + | *The Multi t-test component allows more than two groups to be compared simultaneously. Its shows each Marker Set that has been defined. It will compare all selected sets against all other selected sets. | ||
| + | |||
| + | *A step-down Bonferonni type correction is used to account for multiple testing. | ||
| + | *Results can be viewed in the Volcano Plot and in the Color Mosaic components. | ||
| + | |||
| + | |||
| + | |||
| + | ==References== | ||
| + | t-test [http://www.socialresearchmethods.net/kb/stat_t.htm] | ||
Revision as of 18:02, 19 May 2006
Contents
In this tutorial, you will:
- Get acquainted with the t-Test and Multi t-Test
- Apply a t-Test and Multi t-Test
Before you can continue, geworkbench should be running. Load the data as described in Tutorial - Projects and Data Files.
t-Test
A t-Test analysis can be used to identify markers with statistically significant differential expression between sets of microarrays. The t-test determines, for each marker, if there is a significant difference between the two groups (case and control). To perform this analysis, you must classify the sets, set the analysis parameters and view the results in the visualization components. A detailed description of the t-Test parameters is described in online help.
Classify the Sets
This process has already been described in Tutorial - Data Subsets. Briefly,
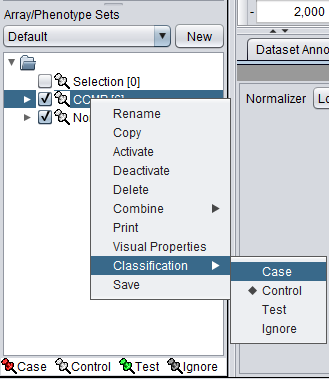
1. Mark the Cardio phenotype a 'Case'. By default, sets are marked as control. Sets classified Case are shown with a red thumbtack icon.
- Right-click on Cardio phenotype.
- Select Classification>Case.
2. Activate the arrays Normal and CCMP by selecting the checkboxes next to the set name.
Set Analysis Parameters
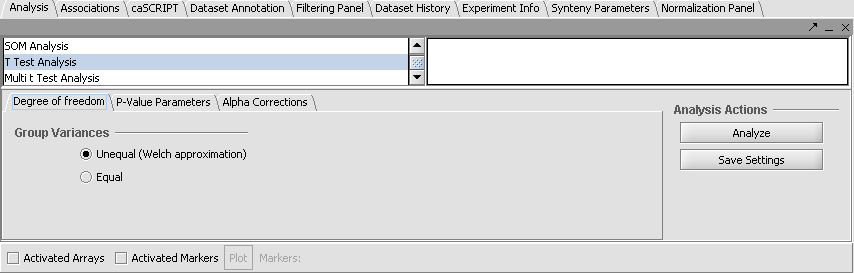
- From the Analysis Panel, select T-Test Analysis.
- Populate the below parameters values and click on Analyze.
- Alpha-corrections tab: Just Alpha.
- P-Value Parameters tab: p-values based on t-distribution. Note that the default alpha (critical p-value) is set to 0.01.
- Degree of Freedom tab: Welch approximation - unequal group variances.
t-Test Results
| Markers which met the significance test are included in a new Marker Set called “Significant Genes”. | 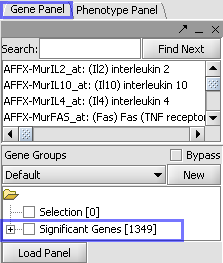
| |
| Ancillary dataset is created in the project window. | 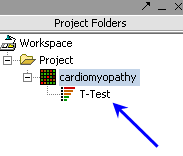
|
The values of the t-Test can be seen in the Color Mosaic panel and the Volcano Plot.
Multi t-test
- The Multi t-test component allows more than two groups to be compared simultaneously. Its shows each Marker Set that has been defined. It will compare all selected sets against all other selected sets.
- A step-down Bonferonni type correction is used to account for multiple testing.
- Results can be viewed in the Volcano Plot and in the Color Mosaic components.
References
t-test [1]