Difference between revisions of "T-test"
(→Outline) |
(→Outline) |
||
| Line 3: | Line 3: | ||
__TOC__ | __TOC__ | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
==Overview== | ==Overview== | ||
Revision as of 12:36, 14 March 2011
Contents
Overview
A t-Test analysis can be used to identify markers with statistically significant differential expression between two sets of microarrays. In geWorkbench, these groups are specified as the "Case" and "Control groups".
There are several steps to setting up a t-test analysis in geWorkbench.
- At least two sets of arrays must be available in the Arrays component.
- At least one each of Case and Control array sets must be activated (by checking the box adjacent to its name).
- At least one set must be classified as "Case". The default classification is "Control".
- The t-test parameters must be set.
After the t-test is run, the results will be displayed graphically, and all markers meeting the significance threshold are placed into a new Marker Set called "Significant Genes".
Preparation
Obtain the file "BCell-100.exp", which is contained in the data/public_data directory of the geWorkbench distribution, or can be directly downloaded from the tutorial data download area.
For tips on loading data files, see Local Data Files and Projects.
t-Test Parameters
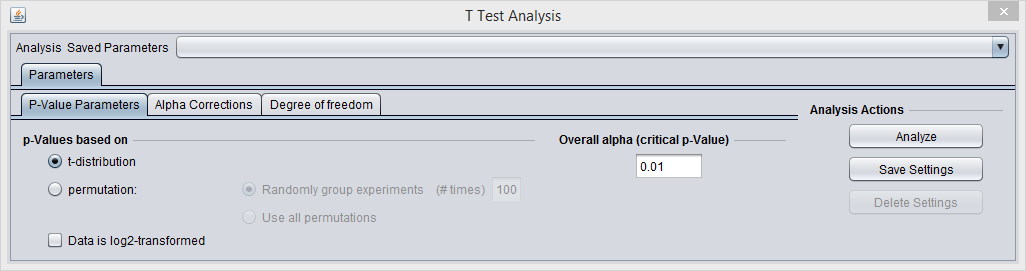
P-value
The p-value can be estimated from 1. the t-statistic (the default) or 2. by permutation.
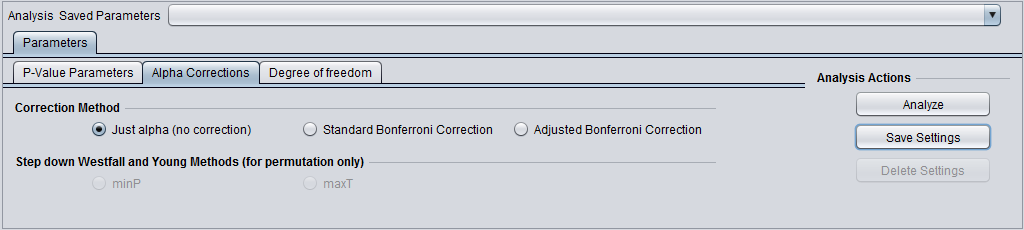
Alpha corrections
For multiple testing (alpha) correction, the following options are offered:
- no correction
- Standard Bonferonni Correction
- Adjusted (step down) Bonferonni Correction.
- Additional methods are available if the p-value is being estimated by permuation:
- minP
- maxT
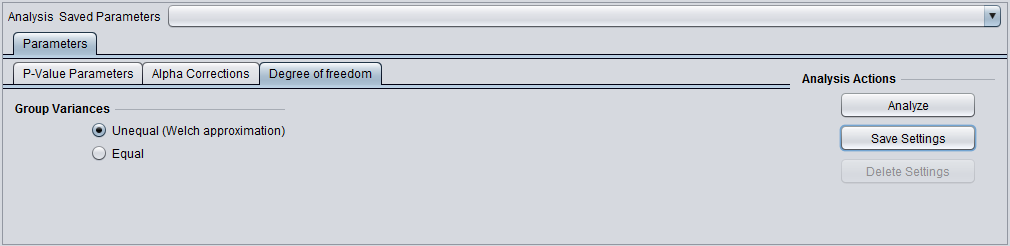
Degrees of Freedom
Group variances can be declared as: 1. unequal (Welch approximation) (the default) 2. Equal.
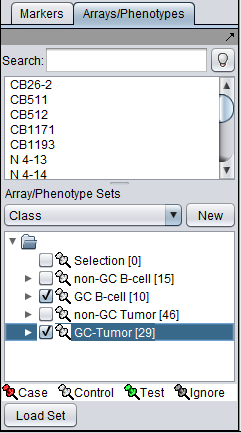
Classification
The desired sets of arrays should be activated in the Arrays/Phenotypes component. This is done by checking the boxes by the desired Sets.
The t-test requires two groups of microarrays to compare. geWorkbench distinguishes the two groups by one being labeled as "Case". By default, all others are considered as control. Note that in the Arrays/Phenotypes component, more than one set of arrays can be marked "Case". All remaining (activated) arrays will then be in the "Control" group.
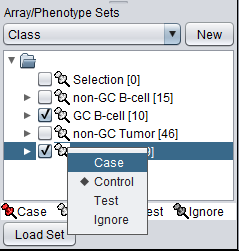
The classification can be made directly by left-clicking on the "thumb-tack" icon adjacent to an array set name.
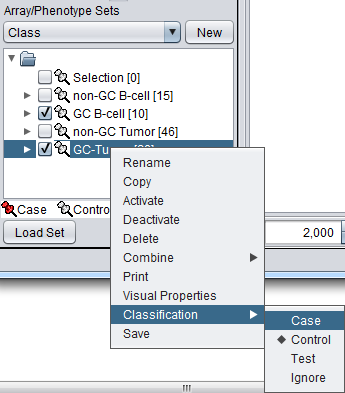
The array classification can also be set by right-clicking on the desired array set and selecting "Classification":
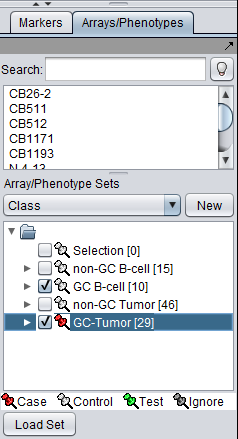
Using either method, the desired array set can be set to be classified as "Case":
The thumbtack image next to activated Array Sets is colored red.
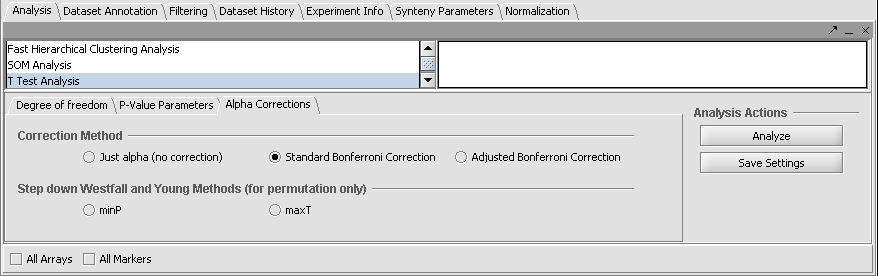
Set Analysis Parameters
- From the Analysis Panel, select T-Test Analysis.
- Various parameters can be adjusted as desired. Here we will use the Standard Bonferonni method, which is the strictest.
- Alpha-corrections tab: Standard Bonferonni.
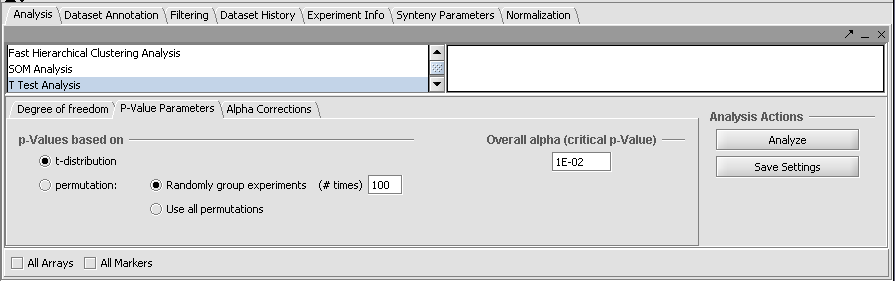
- P-Value Parameters tab: p-values based on t-distribution. Note that the default alpha (critical p-value) is set to 0.01.
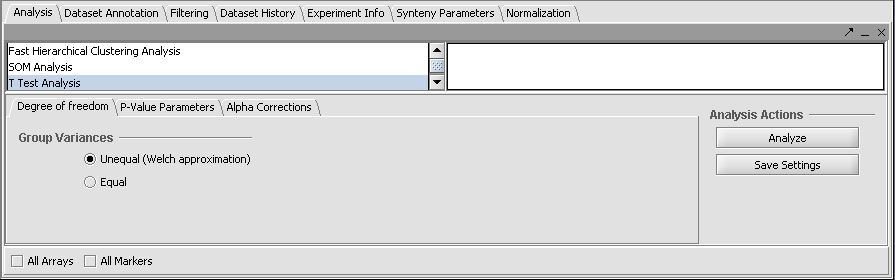
- Degree of Freedom tab: Welch approximation - unequal group variances.
After all the parameters have been set, click Analyze. The results will be returned in three locations: The Project Folder, the Markers component, and the Visualization area.
t-Test Results
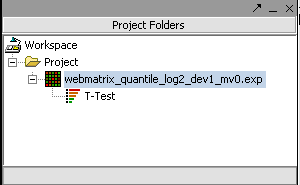
The result is placed into the Projects Folder as a child of the microarray dataset that was analyzed.
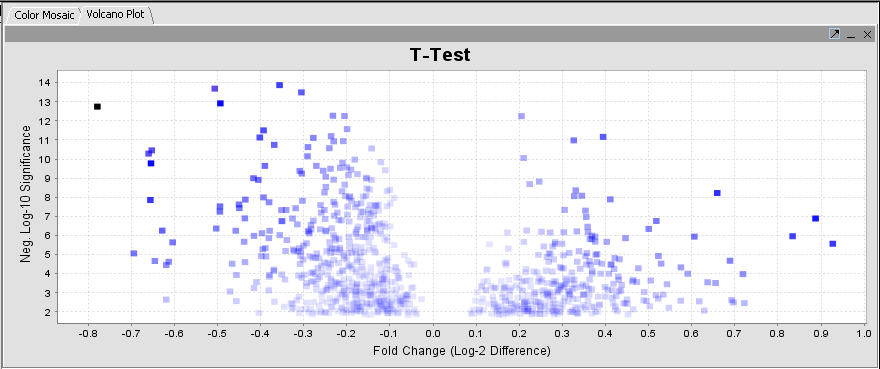
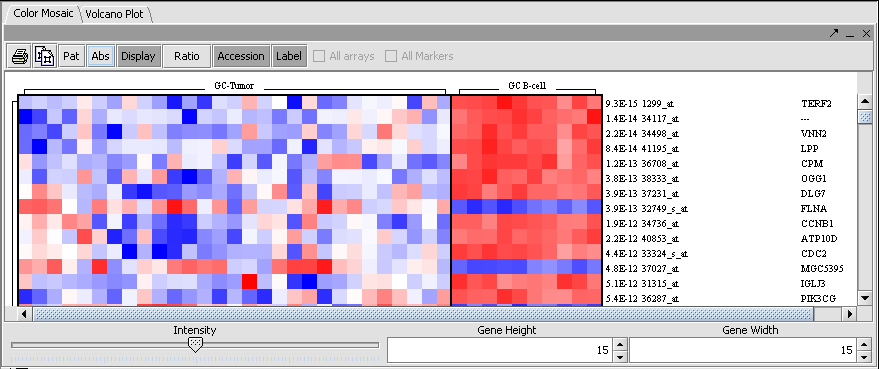
The results are displayed by default using the Volcano Plot visualizer.
The adjacent tab provides a Color Mosaic showing all of the arrays and the p-value calculated for each marker. It also can display annotation for each marker.
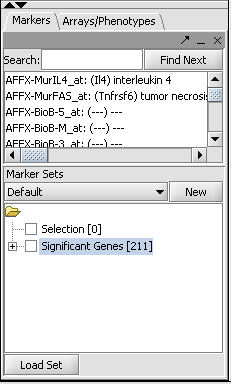
The set of markers which met the minimum signifcance criterion are placed into a new Marker Set labeled "Significant Genes" in the Markers component. The number of markers is shown also.
References
t-test [1]