Difference between revisions of "Promoter Analysis"
(→Overview) |
|||
| Line 4: | Line 4: | ||
=Overview= | =Overview= | ||
| − | The '''Promoter''' component scans one or more sequence | + | The '''Promoter''' component scans one or more sequence profiles against nucleotide sequences that the user has loaded into geWorbench. Motifs from the JASPAR database of transcription factor binding sites are included with the component. Additional motifs can be added by the user. |
The '''Promoter''' component will also display the results of hits found in the '''Pattern Discovery''' component. | The '''Promoter''' component will also display the results of hits found in the '''Pattern Discovery''' component. | ||
| − | |||
| − | |||
=JASPAR CORE database= | =JASPAR CORE database= | ||
Revision as of 16:08, 12 October 2009
|
Home | Quick Start | Basics | Menu Bar | Preferences | Component Configuration Manager | Workspace | Information Panel | Local Data Files | File Formats | caArray | Array Sets | Marker Sets | Microarray Dataset Viewers | Filtering | Normalization | Tutorial Data | geWorkbench-web Tutorials |
Analysis Framework | ANOVA | ARACNe | BLAST | Cellular Networks KnowledgeBase | CeRNA/Hermes Query | Classification (KNN, WV) | Color Mosaic | Consensus Clustering | Cytoscape | Cupid | DeMAND | Expression Value Distribution | Fold-Change | Gene Ontology Term Analysis | Gene Ontology Viewer | GenomeSpace | genSpace | Grid Services | GSEA | Hierarchical Clustering | IDEA | Jmol | K-Means Clustering | LINCS Query | Marker Annotations | MarkUs | Master Regulator Analysis | (MRA-FET Method) | (MRA-MARINa Method) | MatrixREDUCE | MINDy | Pattern Discovery | PCA | Promoter Analysis | Pudge | SAM | Sequence Retriever | SkyBase | SkyLine | SOM | SVM | T-Test | Viper Analysis | Volcano Plot |
Contents
Overview
The Promoter component scans one or more sequence profiles against nucleotide sequences that the user has loaded into geWorbench. Motifs from the JASPAR database of transcription factor binding sites are included with the component. Additional motifs can be added by the user.
The Promoter component will also display the results of hits found in the Pattern Discovery component.
JASPAR CORE database
The motif datafile currently included with geWorkbench is version 3.0 of the JASPAR CORE database (http://jaspar.genereg.net/) . It contains 138 curated, non-redundant profiles. These "profiles are derived from published collections of experimentally defined transcription factor binding sites for multi-cellular eukaryotes. The database represents a curated collection of target sequences" (JASPAR Documentation).
The datafile used "MATRIX_DATA.txt" found at http://jaspar.genereg.net/html/DOWNLOAD/mySQL/JASPAR_CORE_2008/. This file is stored within the Promoter component of geWorkbench.
The profiles are represented by counts of how many times each of the four nucleotide bases occurs at a particular position in the aligned sequences.
Working with the Promoter graphical interface
Layout
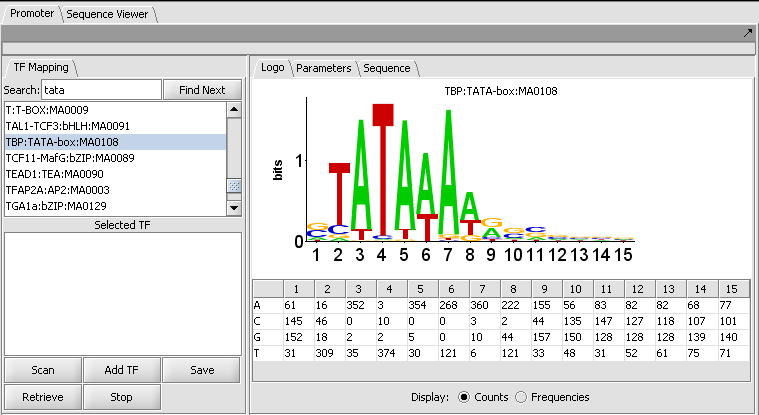
The figure shows the display of profile "TBP:TATA-box:MA0108" from the list of those included in JASPAR.
The LOGO display
The LOGO display implements the method of Schneider to display the information at each position in a motif. Briefly, the total height of the column of letters at a position shows the information available, on a scale of 0 to 2 bits (the information needed to represent the 4 possible nucleotide bases at each position). The relative heights of each letter in a column show their individual contribution to the information at that position.
The LOGO display in geWorkbench implements the "small sample correction" described by Schneider, the magnitude of which depends on the number of sequences aligned to generate the profile. The correction is subtracted from the calculated information content at each position, with a minimum value (floor) of zero being displayed.
The Parameters tab
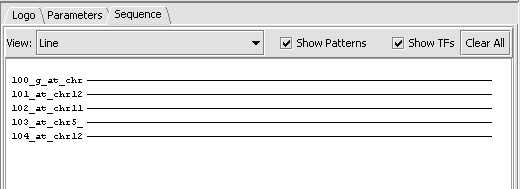
The Sequence tab:
Running and viewing a scan
- Hit Scan.
- Check the box "Show TFs".
- Hits of this motif against the sequences are displayed in the sequence window.
References
Sandelin A, Alkema W, Engstrom P, Wasserman WW, Lenhard B. (2004) JASPAR: an open-access database for eukaryotic transcription factor binding profiles. Nucleic Acids Res. Jan 1;32(Database issue):D91-4 (link to paper).
Schneider TD, Stephens RM. (1990) Sequence logos: a new way to display consensus sequences. Nucleic Acids Res. Oct 25;18(20):6097-100. (link to paper)
Vlieghe D, Sandelin A, De Bleser PJ, Vleminckx K, Wasserman WW, van Roy F, Lenhard B. (2006) A new generation of JASPAR, the open-access repository for transcription factor binding site profiles. Nucleic Acids Res. Jan 1;34(Database issue):D95-7.