ARACNe
Contents
- 1 Overview
- 2 Setting up an ARACNe run
- 3 Services (Local vs Grid)
- 4 Viewing ARACNe results
- 5 Dataset History
- 6 Example of running ARACNe
- 7 Technical Notes
- 8 References
Overview
ARACNe (Algorithm for the Reconstruction of Accurate Cellular Networks) (Basso et al. 2005; Margolin, Nemenman et al. 2006) is an information-theoretic algorithm used to identify transcriptional interactions between gene products using microarray expression profile data. The resulting network is displayed using the Cytoscape component. ARACNe can be used to predict potential functional associations among genes, or to predict novel functions for uncharacterized genes, by identifying statistical dependencies between genes. The results take the form of a matrix of candidate interactions, also called an adjacency matrix, which can be used for further network visualization and analysis. ARACNe has been used to reconstruct networks in mammalian cells through appropriate choice of dataset.
ARACNe performs best with a dataset containing data from a minimum of 100 microarrays (see Margolin, Wang et al. 2006) and representing a number of different states of the same cellular system - for example, cells lines of varying phenotype, or cells subjected to a variety of experimental perturbations. Initial work with ARACNe was performed using a large collection (about 340) of B-cell lines of various phenotypes (Basso et al. 2005). For the "Adaptive Partitioning" option, there is no upper limit on the number of arrays that can be included. For the "Fixed Bandwidth" option, arrays in excess of 300 can lead to long computation times. A subset of the B-Cell dataset, derived from 100 arrays, is included with geWorkbench (Bcell-100.exp).
ARACNe can perform two separate calculations:
- Mutual Information: The mutual information (MI) of one or more marker's expression profile(s) is calculated against all other active markers.
- Data Processing Inequality (DPI): The DPI calculation (triangle inequality) is used to remove the weakest interaction (edge) between any three markers. That is, if a MI value is available between each of three possible pairings of three markers, the weakest interaction of the three will be removed from the output. This has the intent of removing indirect interactions. For example, if A->B->C, the interaction A->C will likely be weaker than A->B or B->C and would be removed. A tolerance can be set which relaxes this screening to account for uncertainty in the MI calculation.
Parameters described below allow one to incorporate a list of putative transcription factors and optimize the run to discover targets that they may regulate.
Current versions of geWorkbench (starting with release 1.7.0) use a new implementation of ARACNe, ARACNe2, that supports two important new features:
- Algorithm - A new algorithm, termed "Adaptive Partitoning", which is much faster than the original "fixed bandwidth" method was added.
- Mode - The user can choose to custom-calculate optimal run parameters for a given dataset.
For a dataset with a simple, monotonic relationship between input and output, analysis with a normal (e.g. Pearson's) correlation function may be the most suitable method. Mutual information does not require a monotonic relationship. Where the input/output function is non-linear or irregular, a mutual information method such as ARACNe may be able to find relationships that correlation will not find.
Further information on ARACNe is available in the References section below.
Setting up an ARACNe run
Prerequisites
- To use the ARACNe routine, first check that it has been loaded in the Component Configuration Manager.
- ARACNe is found in the list of available analysis routines in the lower-right Commands quadrant of geWorkbench.
- A microarray dataset of sufficient size and phenotypic diversity is needed (See the Overview, above).
- Load the microarray dataset into the Project Folders component. If available, associate a gene annotation file with the dataset. This will allow the results to be displayed in consolidated fashion in Cytoscape by gene rather than by marker (individual probeset) name.
Parameters and Settings
Algorithm
Two algorithms are offered with which to calculate the pairwise mutual information between markers:
- Adaptive Partitioning (default) - should generally be used for all new calculations.
- Fixed Bandwidth - previous, slower algorithm using a fixed Gaussian kernel.
Adaptive Partitioning
Adaptive Partitioning was added with the incorporation of the ARACNe2 code into geWorkbench ( as of version 1.7.0). Adaptive Partitioning is much faster than the original Fixed Bandwidth method, and is also considered to produce superior results under certain circumstances. Unlike the Fixed Bandwidth method, it does not used a fixed kernel-width when calculating the MI.
Fixed Bandwidth
Fixed Bandwidth was the original algorithm used in ARACNe and is included for compatibility with previous releases. This method uses a kernel-width parameter for a Gaussian function used to calculate the MI.
Mode
Used to control the calculation and use of runtime parameters from the input dataset.
- PREPROCESSING - calculates the required parameters and writes them to parameter files.
- DISCOVERY - The ARACNe mutual information calculation is run. Uses pre-calculated parameter files as needed if they are present.
- COMPLETE - Preprocessing and Discovery runs are combined into a single step.
PREPROCESSING
In this mode, runtime parameters are calculated, but no MI calculation is performed. Preprocessing for a given combination of dataset and algorithm need be run only once. The results are written to one or two files in the geWorkbench root directory. The names used for these files incorporate both the name of the dataset and the name of the algorithm, and thus are specific to the particular combination. Each time ARACNe is run in Discovery mode, it will look for the dataset-specific parameter files in its root directory. If the files are not found (Preprocessing has not been run), default parameter values will be used.
- Fixed Bandwidth (FBW) algorithm - two files are written to the geWorkbench root directory, one containing parameters for calculating the kernel width, and the other containing parameters for calculating a MI threshold from a specified P-value.
- Adaptive Partitioning (AP) algorithm - only the parameter file for calculating a MI threshold from a specified P-value is written. No kernel-width parameter is used.
Preprocessing files included with geWorkbench
Preprocessing as described above was run on the Bcell-100.exp dataset included with geWorkbench. The resulting ARACNe parameter files are also included in the geWorkbench root directory. They will be used by default when the Bcell-100.exp dataset is used in tutorials. Note that if you rerun the preprocessing step, the relevant file(s) will be overwritten.
The parameter files included in geWorkbench are:
- Bcell-100.exp_ARACNe_AP_threshold.txt - Adaptive Partitioning, Pvalue-to-MI threshold conversion parameters.
- Bcell-100.exp_ARACNe_FBW_kernel.txt - Fixed Bandwidth, kernel width calculation parameters.
- Bcell-100.exp_ARACNe_FBW_threshold.txt - Fixed Bandwidth, Pvalue-to-MI threshold conversion parameters.
DISCOVERY
The ARACNe mutual information and the DPI (if selected) calculations are run. If dataset-specific parameter files are present, they will be used as needed (based on settings selected for Kernel Width and Threshold).
COMPLETE
A preprocessing run will be performed followed immediately by a Discovery run. The dataset-specific parameter files created during the Preprocssing step will be used if needed (based on settings selected for Kernel Width and Threshold).
When is preprocessing not needed?
The preprocessing step can be time consuming. If you are for example using Adaptive Partioning, and decide you do not need to specify a p-value threshold for accepting edges, then you can just set a MI value as the threshold and proceed directly to Discovery mode. This will however make interpreting results more difficult.
If ARACNe does not find the dataset-specific parameter files it needs as described above, it will use by default parameters calculated from the B-cell dataset (see Margolin et al., 2006).
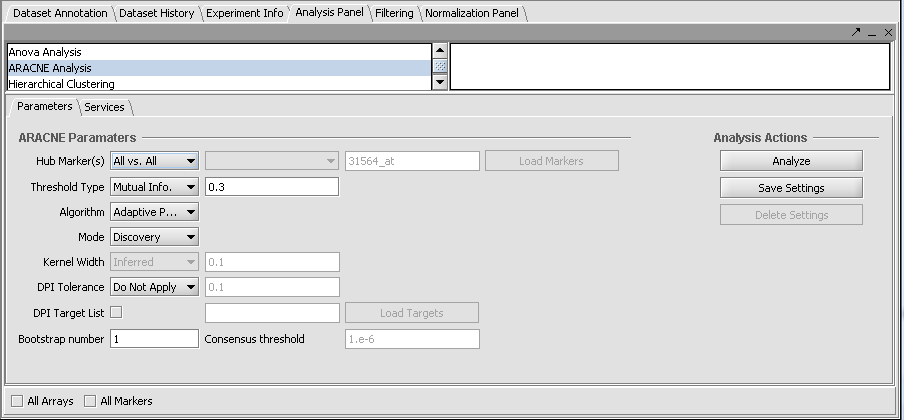
Hub Marker(s)
Specifies which gene markers will be treated as "hubs" in the ARACNE mutual information (MI) calculation. The mutual information is calculated for each specified hub marker against all other markers in the submitted dataset. For many uses, it is suggested to use a defined list of known transcription factors as hub markers, rather than using the "All-vs-All" setting.
- All vs All - The MI of every pair of markers in the dataset is computed, that is, each is used as a hub.
- From Sets - allows a set of markers defined in the Markers component to be chosen from a pulldown menu. Alternatively, the user can type in the names of desired markers directly as a comma separated list.
- From File - allows a comma-separated list of markers to be read in from a file by clicking Load Markers..
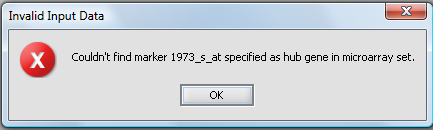
Hub marker(s) must appear in active marker set
If a set of markers is activated in the Markers component, rather than using all markers, then the chosen hub marker(s)must also be included in an active set. If the hub marker is missing from the active sets, then an error dialog will be displayed. In the below picture, the marker 1973_s_at was entered into the hub field without being part of a subset of markers that had been activated:
Threshold Type
This drop-down specifies the type of threshold to be used and can take the values “Mutual Info” or “P-value”. The actual value entered into the adjacent text area is always a number between 0 and 1.
Kernel width
The Kernel width is a scaling parameter used for fitting a Gaussian function to the data when running the FIXED_BANDWIDTH algorithm only, otherwise this field is disabled. If used, the value can be either inferred or specified directly.
- Inferred: If PREPROCESSING has been run on the dataset (mode is set to PREPROCESSING or COMPLETE), the kernel width is calculated directly from those results. If PREPROCESSING has not been run, the kernel width is inferred based on parameters fitted to a large B-cell dataset (Margolin et al, 2006), extrapolated for the number of samples in the dataset being tested.
- Specify: The user can enter a value for the kernel width directly, e.g. based on a prior calculation with this dataset.
DPI Tolerance
The Data Processing Inequality (triangle inequality)can be used to remove the effects of indirect interactions, e.g. if TF1->TF2->Target, DPI can be used to remove the indirect action of TF1 on the target. Stated another way, the DPI can be used to remove the weakest interaction of those between any three markers. The DPI tolerance specifies the degree of sampling error to be accepted, as with a finite sample size an exact value MI can not be calculated. The higher the tolerance specified, the fewer the edges that will be removed.
- Do Not Apply - Do not use the DPI.
- Apply - DPI is applied using the threshold value (between 0.0 and 1.0) specified in the adjacent text field. The higher the threshold, the weaker the screening and the more edges will be included in the final output.
DPI Target List
The DPI target list can be used to limit the ARACNE calculation to transcriptional networks. It is used to screen out spurious regulatory interaction signals of genes that are tightly co-expressed but are not in a regulatory relationship to each other, for example genes for two proteins that are in a physical complex and hence always produced in the same amounts. A comma-separated list can be typed in, or it can be loaded from an external file. If used, the DPI Target List should contain all markers that are annotated as transcription factors. Signaling proteins could also be included.
- Details: If the box is checked, the user selects and loads a file which specifies markers (which should be a list of one or more presumptive transcription factors) which will be given preferential treatment during the DPI edge-removal step. Edges originating from markers on this list will not be removed by edges originating from markers not on this list. However, for DPI calculations where all three markers are members of the list, the weakest connecting edge may still be removed.
Bootstrapping
Bootstrap analysis can be used to generate a more reliable estimate of statistical significance for the interactions. Please see Margolin et al. 2006, Nature Protocols, Vol 1, No. 2, pg. 663-672 for further details (full reference below). Briefly, repeated runs of ARACNE are made, with arrays drawn at random from the full dataset with replacement. The same number of arrays is drawn each time as is present in the original dataset. A permutation test is then used to obtain a null distribution, against which the statistical significance of support for each network edge connection (interaction) can be measured.
- Bootstrap number: Specifies the number of bootstrapping runs to perform.
- Consensus threshold (for bootstrapping only): After the bootstrapping runs are made, a permutation test is used to estimate the significance of interactions. The consensus threshold sets the cutoff point for calling the interactions significant and returning them in the final adjacency matrix
Array and Marker Set Overrides
- All Markers: checking this box overrides any activated marker set in the Markers component.
- All Arrays: checking this box overrides any activated array set in the Arrays/Phenotypes component.
Analysis Actions
- Analyze - start the ARACNe analysis
- Save Settings, Delete Settings - The geWorkbench analysis framework provides a standard method for saving one or more different sets of parameter settings per each type of analysis component. Please see the Analysis Framework chapter for further details.
Services (Local vs Grid)
ARACNe can be run either locally within geWorkbench, or remotely as a grid job on caGrid. See the Grid Services section for further details on setting up a grid job.
Special Note on running in PREPROCESSING mode on caGRID
When ARACNew is run in PREPROCESSING mode on a grid node, it writes the parameter files to its execution directory on the grid node and exits. No file is returned to geWorkbench. As currently implemented, the ARACNe server detects the lack of a file to return (normally it returns an adjacency matrix) and reports an error. This error can simply be ignored. If ARACNe2 is run in COMPLETE or DISCOVERY mode this error will not occur because both return adjacency matrices.
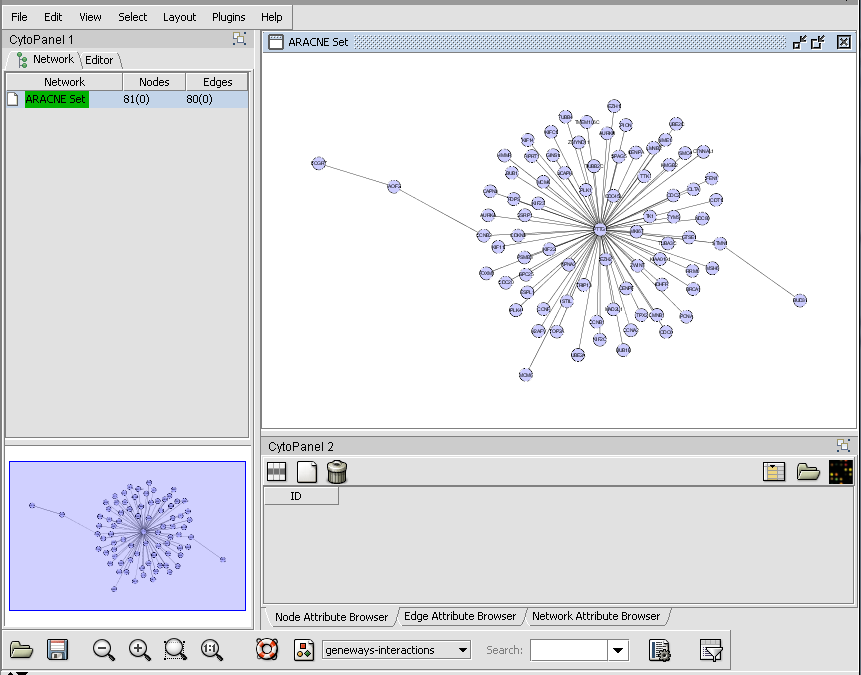
Viewing ARACNe results
- The result of an ARACNe run is an "adjacency matrix". it contains the mutual information value for each pair of markers which exceeded the specified MI threshold. The adjacency matrix is placed into the Project Folders component as a child of the dataset that was analyzed.
- The adjacency matrix can be visualized automatically in the Cytoscape component, as shown below.
The integration between Cytoscape and geWorkbench allows for two-way interactions between them:
- Nodes selected in Cytoscape appear in the Marker Sets component in the set "Cytoscape selection".
- Any set of markers in the Marker Sets component can be projected onto the Cytoscape display, which will cause any matching nodes there to be highlighted.
This interaction is demonstrated further on the Cytoscape tutorial page.
The Cytoscape Viewer maintains a list of networks which it has currently loaded. It allows individual loaded networks to be deleted. However, the network can be reloaded by clicking on its entry in the Project Folders component. Cytoscape controls are more fully described in the Cytoscape component tutorial.
Dataset History
Details about each run are maintained in the Dataset History component. With the ARACNe result node highlighted in the Project Folders component, the Dataset History display includes the following information:
- Input file name
- Output file name
- Algorithm
- Mode
- No. bins
- MI threshold
- MI threshold calculated from P-Value - If supplied, the p-value used to set the MI threshold.
- DPI tolerance
- Hub markers
- A listing of the microarrays used.
- A listing of the markers used.
Example of running ARACNe
This example uses the Bcell-100.exp dataset available in the data/public_data directory of geWorkbench, and further described on the Download page. Briefly, this dataset is composed of 100 Affymetrix HG-U95Av2 arrays on which various B-cell lines, both normal and cancerous, were analyzed. Thus it explores a potentially wide variety of expression phenotypes.
Prerequisites
- (Optional) Obtain the annotation file for the HG-U95Av2 array type from the Affymetrix NetAffx website (http://www.affymetrix.com/analysis/index.affx). The name will be similar to "HG_U95Av2.na29.annot.csv", where na29 is the version number. Loading the annotation file associates gene names and other information with the Affymetrix probeset IDs (see the geWorkbench FAQ for details on obtaining these files).
Loading the example data
- Load the Bcell-100.exp dataset into geWorkbench as type "Affymetrix File Matrix". (See Local Data Files).
- When prompted, and if desired, load the annotation file.
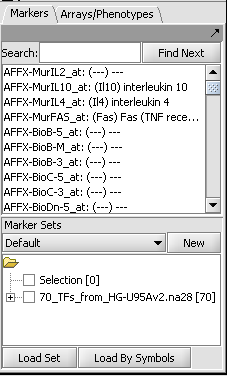
- In the Marker Sets component, load the file "70_TFs_from_HG-U95Av2.na28.csv" (Load Set button). This file is included in the geWorkbench tutorial data.
Setting up the parameters and starting ARACNe
1. In the geWorkbench commands area, select the "ARACNe" analysis.
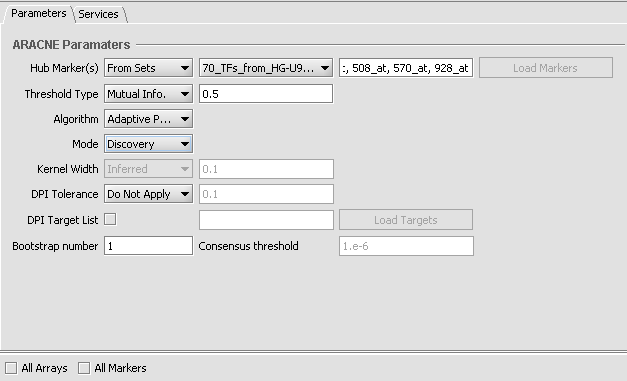
2. Set up the ARACNe parameters as shown
- Hub Markers: From Sets - Choose the set of 70 loaded above "70_TFs_from_HG-U95Av2.na28".
- Threshold: Mutual Info - Set value of 0.5.
- Algorithm: Adaptive Partitioning.
- Mode: Discovery
- DPI Tolerance: Do Not Apply
3. Press the "Analyze" button to launch the job. On a current generation desktop machine expect this example to run for several minutes.
4. The resulting network is the one depicted above in the "Viewing ARACNe results" section.
Technical Notes
- The set of hub markers, if supplied, is the same as using the "-s" subnet option with the standalone ARACNe executable. "... a list of probes for which a subnetwork will be constructed".
- The DPI target list, if supplied, is the same as using the "-l" transcription factor option with the standalone ARACNe executable. "...a list of probes annotated as transcription factors in the input dataset".
- "This option is ideal for transcriptional network reconstruction. If provided, DPI will not
remove any connection of a transcription factor (TF) by connections between two probes not annotated as TFs. This option is often used in conjunction with '-s', which specifies a list of probes that are either the same or a subset of the probes specified by '-l'".
References
- Basso K, Margolin AA, Stolovitzky G, Klein U, Dalla-Favera R, Califano A: Reverse engineering of regulatory networks in human B cells. Nat Genet 2005, 37(4):382-390 (link to paper).
- Margolin AA, Nemenman I, Basso K, Wiggins C, Stolovitzky G, Dalla Favera R and Califano A, (2006) ARACNE: An Algorithm for the Reconstruction of Gene Regulatory Networks in a Mammalian Cellular Context, BMC Bioinformatics;7(Suppl.1):S7 (link to paper)
- Margolin, A., Wang, K., Lim, W.K., Kustagi, M., Nemenman, I., and Califano, A. Reverse Engineering Cellular Networks. Nature Protocols (2006), Vol 1, No. 2, pgs. 663-672 (link to paper)