MatrixREDUCE
Contents
Outline
This tutorial contains
- . an overview of MatrixREDUCE and
- . instructions....
Overview
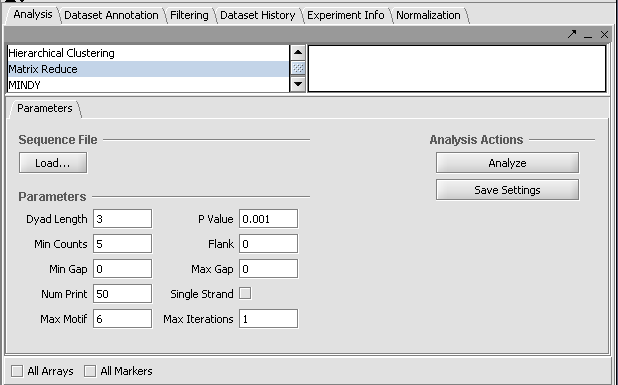
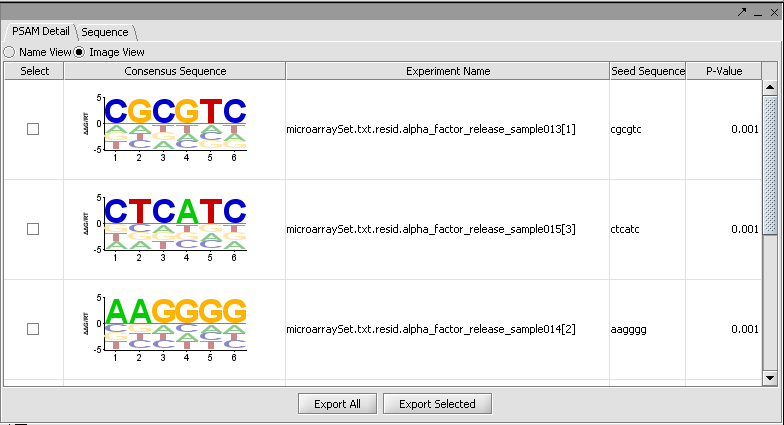
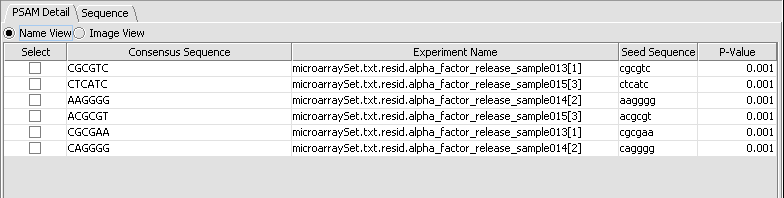
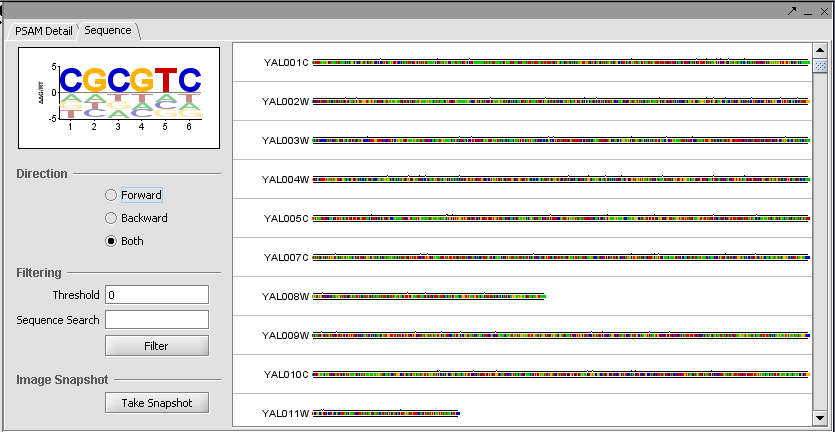
MatrixREDUCE is a tool for inferring cis-regulatory elements and transcriptional module activities from microarray data. The run parameters are entered in the Analysis section of the application and the results are displayed in the visualization pane.
Data Files
MatrixREDUCE operates on two input files: a microarray data set and a FASTA file containing the DNA sequences corresponding to the regulatory region of genes probed in the microarray set. The gene/probe identifiers used in the microarray dataset and the sequence identifiers used in the FASTA file must match. However, case is not important; the program will change the identifiers to lower case before attempting to match the records. If an identifier appears more than once in a file, the last instance is used.
Example
In this example, we will use two files that are included in the data directory of the geWorkbench distribution iteself. They are SpellmanReduced.txt and Y5_600_Bst.fa. SpellmanReduced.txt contains a subset of the data from Spellman XXXXX. Y5_600_Bst.fa contains the corresponding upstream DNA sequences for these genes.