Difference between revisions of "Filtering"
| Line 1: | Line 1: | ||
| + | {{TutorialsTopNav}} | ||
| + | |||
| + | |||
| + | |||
| + | ==Overview== | ||
| + | |||
| + | Filtering can be used to remove low quality data or reduce the size of the dataset by removing less interesting data. Most geWorkbench filters allow the user to specify a minimum number or percentage of arrays that must meet that filter's critereon before the marker will be removed. | ||
| + | |||
| + | ==Filter Configuration== | ||
| + | |||
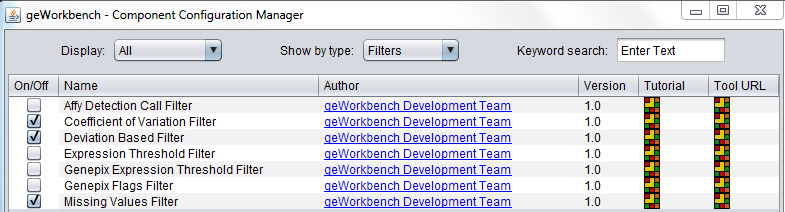
| + | Some filters are not loaded by default in geWorkbench. To configure these filters, use the Component Configuration Manager (Tools->Component Configuration). | ||
| + | |||
| + | [[Image:Filters_in_CCM.png]] | ||
| + | |||
| + | |||
| + | ==Available Filters== | ||
| + | |||
| + | {|style="border: 1px solid lightGray" | ||
| + | !Filter||Description | ||
| + | |- | ||
| + | |- | ||
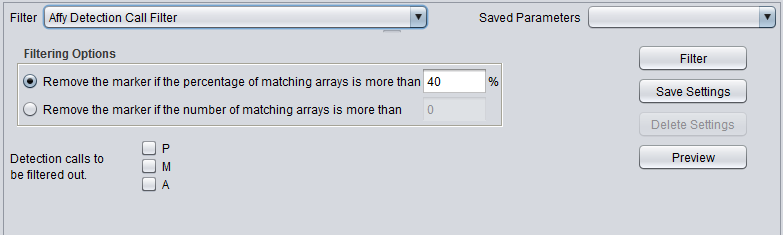
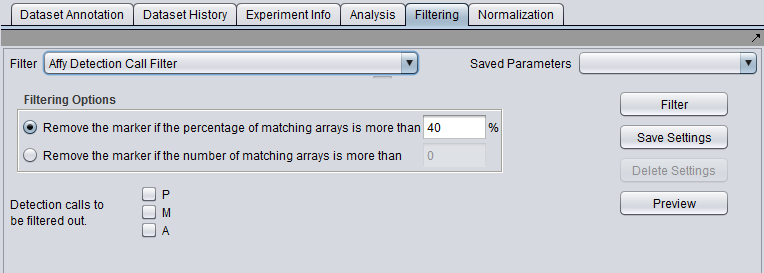
| + | |'''Affy Detection Call''' ||Applicable to Affymetrix data only. Filter on Present, Marginal or Absent calls. | ||
| + | |- | ||
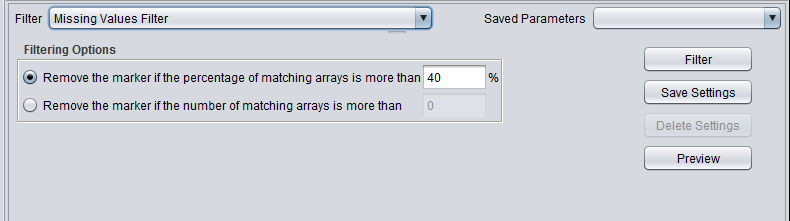
| + | |'''Missing values''' ||Removes markers that have “missing” measurements in more than the specified number (or percentage) of microarrays. | ||
| + | |- | ||
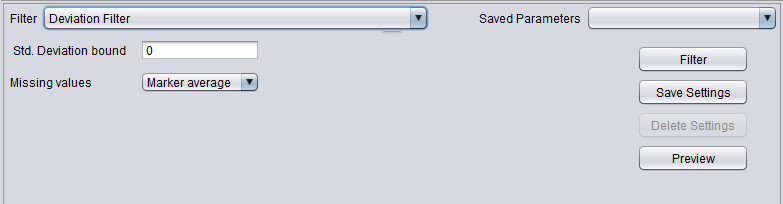
| + | |'''Deviation''' ||Removes markers whose '''standard deviation''' is less than the specified value across all microarrays. | ||
| + | |- | ||
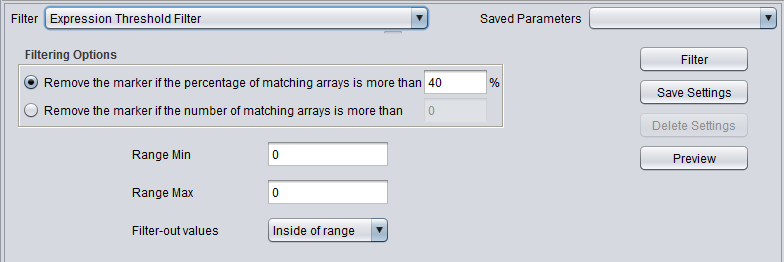
| + | |'''Expression Threshold''' ||Removes markers where more than a specified number (or percentage) have values inside (or outside) a user-defined range. | ||
| + | |- | ||
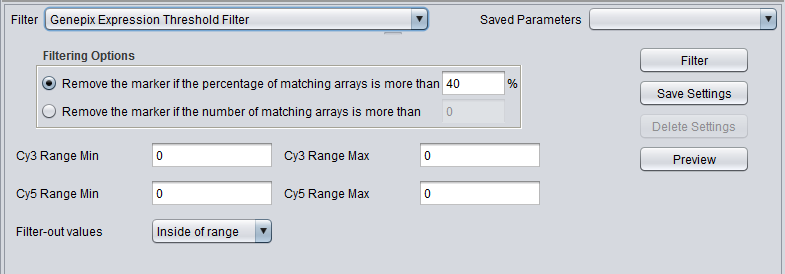
| + | |'''Genepix Expression Threshold Filter'''||Applicable to 2-channel arrays (Genepix) data only. Defines applicable ranges for each channel, and removes markers for which for more than a specified number (or percentage) of markers either channel intensity is inside (or outside) the defined range. | ||
| + | |- | ||
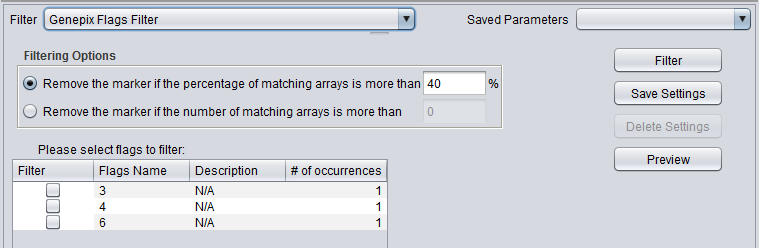
| + | |'''GenePix Flags''' ||Remove markers where more than a specified number of values match the selected flag (Flagged in GenePix software). | ||
| + | |- | ||
| + | |} | ||
| + | |||
| + | |||
| + | |||
[[Image:Filtering-Affy_detection_call.png]] | [[Image:Filtering-Affy_detection_call.png]] | ||
| Line 13: | Line 50: | ||
[[Image:Filtering-Missing_Values_Filter.png]] | [[Image:Filtering-Missing_Values_Filter.png]] | ||
| − | |||
| − | |||
Revision as of 16:29, 4 June 2010
Overview
Filtering can be used to remove low quality data or reduce the size of the dataset by removing less interesting data. Most geWorkbench filters allow the user to specify a minimum number or percentage of arrays that must meet that filter's critereon before the marker will be removed.
Filter Configuration
Some filters are not loaded by default in geWorkbench. To configure these filters, use the Component Configuration Manager (Tools->Component Configuration).
Available Filters
| Filter | Description |
|---|---|
| Affy Detection Call | Applicable to Affymetrix data only. Filter on Present, Marginal or Absent calls. |
| Missing values | Removes markers that have “missing” measurements in more than the specified number (or percentage) of microarrays. |
| Deviation | Removes markers whose standard deviation is less than the specified value across all microarrays. |
| Expression Threshold | Removes markers where more than a specified number (or percentage) have values inside (or outside) a user-defined range. |
| Genepix Expression Threshold Filter | Applicable to 2-channel arrays (Genepix) data only. Defines applicable ranges for each channel, and removes markers for which for more than a specified number (or percentage) of markers either channel intensity is inside (or outside) the defined range. |
| GenePix Flags | Remove markers where more than a specified number of values match the selected flag (Flagged in GenePix software). |