Difference between revisions of "LINCS Query"
(→LINCS Query and Display) |
(→LINCS Query and Display) |
||
| Line 24: | Line 24: | ||
** The match can occur anywhere in the drug name. | ** The match can occur anywhere in the drug name. | ||
| − | * '''Max Results''' - | + | * '''Max Results''' - If the box is checked, return and displays only the top ''n'' results by the value shown in the "Score" column, where ''n'' is the value entered into the adjacent text field. |
==Experimental vs Computational== | ==Experimental vs Computational== | ||
Revision as of 16:58, 19 June 2013
Contents
Overview
LINCS - Library of Integrated Network-based Cellular Signatures
This component provides for query and display of data generated by the Columbia LINCS Technology U01 and Computation U01 Centers. It provides experimental and computational results for drug mode of action and similarity calculations, and for synergy experiments.
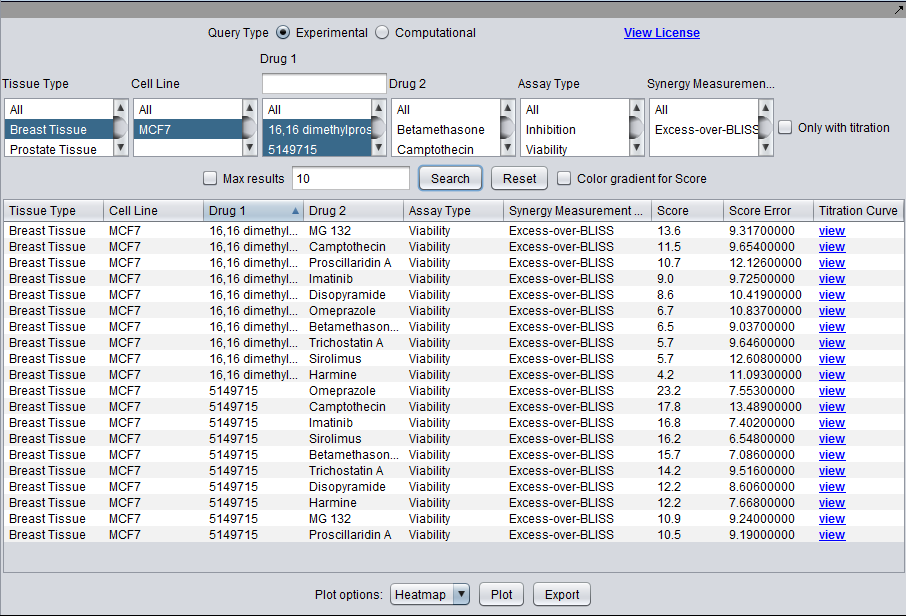
LINCS Query and Display
Query for experimental synergy results for tested drug pairs, or computational drug similarity results based on gene expression experiments. In the current experimental design, drug 1 and drug 2 not symmetric, they are generally drawn from separate lists of compounds. This difference is preserved in the database.
- Tissue Type - The tissue from which the tested cell line was originally derived. More than one tissue can be chosen for a single query.
- Cell Line - The cell line tested in a particular experiment. More than one cell line can be chosen for a single query (multiple selection), subject to any constraint on "Tissue Type".
- Drug 1 - Entries stored as "drug 1" in a pair, subject to previous choices of tissue type or cell line, if any. One or more drugs (multiple selection) must be chosen.
- Drug 2 - Entries stored as "drug 2" in a pair, given current constraints on tissue type, cell line, and drug 1. More than one drug can be chosen for a single query (multiple selection).
- Dynamic Search on Drug 1 - currently the number of drugs in "Drug 1" can be considerably larger than in "Drug 2". For this reason, a dynamic search box is offered above the "Drug 1" selection. As each character is typed, the "Drug 1" list will be updated to only show matching drugs.
- The match is not case-sensitive.
- The match can occur anywhere in the drug name.
- Max Results - If the box is checked, return and displays only the top n results by the value shown in the "Score" column, where n is the value entered into the adjacent text field.
Experimental vs Computational
Radio buttons are used to select whether to query for experimental or computational results. Only one type can be chosen at a time. These result types are held in separate tables in the database and the scores have different ranges.
The controls specific to each type are:
Experimental
- Assay Type (phenotypic readout): Viability, proliferation, apoptosis, cell morphology, oxidization state, etc.
- Synergy Measure (Excess over Bliss, Excess over highest single agent, Combination Index, etc.).
Computational
- Similarity Algorithm (MARINa (GSEA-2), IDEA, DEMAND, combined, etc.). This entry will combine the algorithm used with the distance method used, e.g. GSEA-2, in the case where an algorithm has more than one possible measure.