MarkUs
Contents
Overview
MarkUs is a web server to assist the assessment of the biochemical function for a given protein structure. MarkUs identifies related protein structures and sequences, detects protein cavities, and calculates the surface electrostatic potentials and amino acid conservation profile. The results can be browsed by an interactive web interface that allows one to integrate Gene Ontology terms, UniProt features, and the Enzyme Classification. The MarkUs website is at http://luna.bioc.columbia.edu/honiglab/mark-us/cgi-bin/submit.pl.
MarkUs is available as a local or grid service in geWorkbench. Details about running grid services are available in the Grid Services tutorial.
MarkUs Server Documentation
Direct Submission of a job to the MarkUs server
http://luna.bioc.columbia.edu/honiglab/mark-us/cgi-bin/submit.pl
This page contains a concise description of the goals of the MarkUs server and a link to the direct web job submission page.
MarkUs Server NESG tutorial
http://luna.bioc.columbia.edu/honiglab/nesg/documentation/tutorial.html
Step by step illustration of submitting a job and interpreting the results.
Detailed Overview of MarkUs Analysis
http://luna.bioc.columbia.edu/honiglab/nesg/documentation/index.html
Describes each aspect of the Markus analysis results and their presentation.
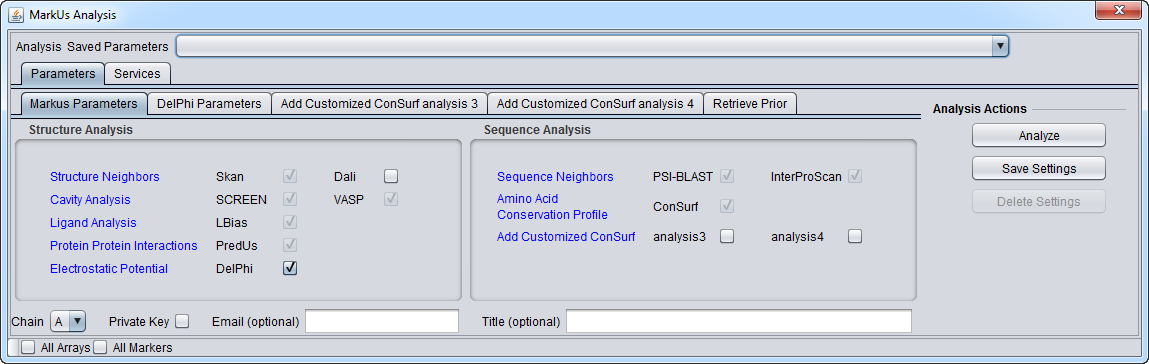
Parameters
Main
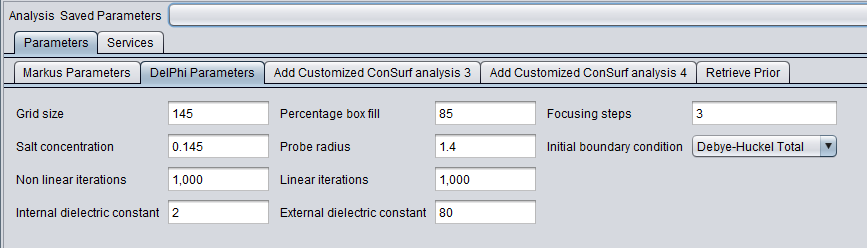
DelPhi
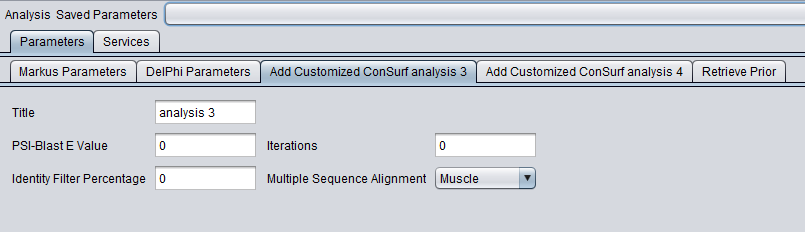
ConSurf
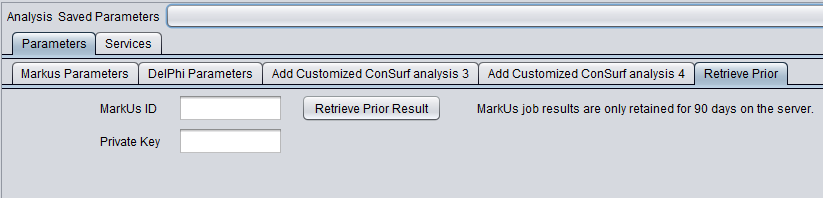
Retrieve Prior Jobs
The results of previously run MarkUs jobs may be retrieved from the MarkUs server at a later time. However, note that results may be deleted after 90 days.
To retrieve prior results, you must have, when the job was run, made a note of the job id and, if requested, the private key.
- MarkUs ID - When a MarkUs job is submitted, a job ID is automatically generated by the MarkUs server. This ID is displayed in the results page, and is also used as the results node name in the Project Folders component.
- Private Key - If the "Private Key" box was checked on the Parameters tab when the job was submitted, the system also generated and displayed this key in the name of the result node. In this case, you must also supply this private key to retrieve the job results.
Results
Private Key
The figure below depicts three separate runs of MarkUs. In the first two, the "Private Key" checkbox was checked in the Parameters tab. The result node name in this case, e.g. "MUS2743&key=QZ860109" contains both the job name, e.g. MUS2743, as well as the private key following the label "key=", in this case "QZ860109".
For jobs which requested a private key, you will need to supply that key in the "Retrieve Prior" tab if you wish to view the results in another invocation of geWorkbench.
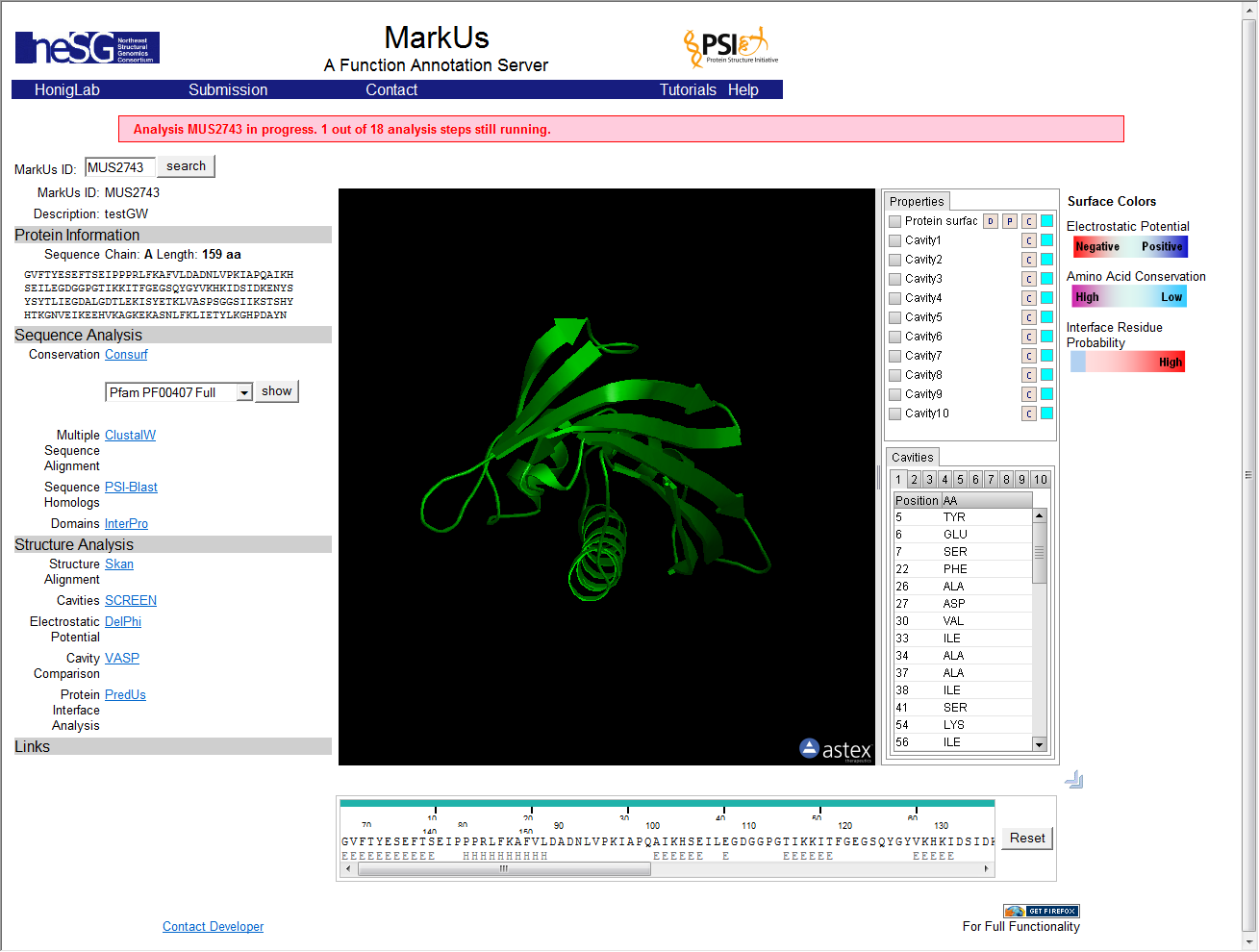
Result Viewer
The result of the analysis is returned to a web-browser window displayed within geWorkbench. The run may take a considerable amount of time to finish, and partial results will be displayed. The window does not auto-refresh. To see the most recent results, right-click on the result display window and select "refresh".
Running MarkUs
The starting point for a MarkUs analysis is a PDB protein structure file.
1. If the MarkUs component has not been loaded in the Component Configuration Manager (CCM), first do so.
2. Load a PDB file for the protein whose structure you would like to analyze.
3. Set the desired parameters. Note that some analyses have check-boxes next to them which are grayed-out. These analyses are always run as part of a MarkUs job and cannot be left out.
4. If the "private key" checkbox is checked, MarkUs will generate a private key which will be displayed in the result node in the Project Folders compnent.
5. Inspect and adjust if desired the parameters to DelPhi. DelPhi calculates the electrostatic field around the molecule.
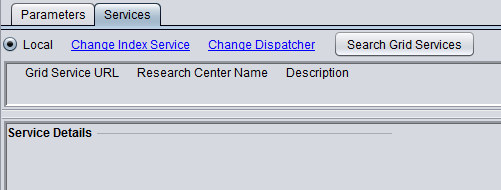
6. MarkUs is available in geWorkbench as a local or grid service. Both local and grid job submission are asynchronous and to the same server, so there is no advantage to using the grid service.
The figure below shows the local service as selected.
7. Return to the Parameters tab.
8. Push the "Analyze" button.
9. A result node will be created in the Project Folders component.
10. If the "Private Key" checkbox was checked, copy down the private key from the new result node in the Project Folders component.
Services (Grid)
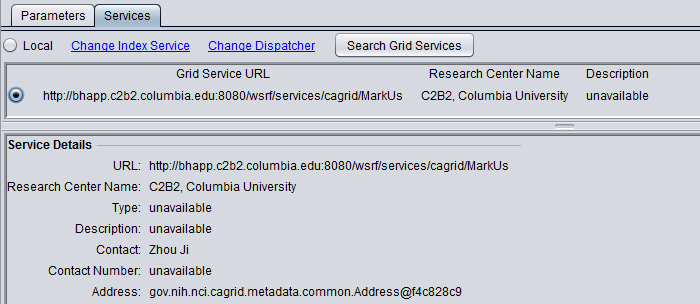
To select the grid service for MarkUs,
- Click on the Services tab.
- Push the "Grid Services" button. This will retrieve available MarkUs grid services from an index server.
- Select the radio button for the desired grid service.
See the Grid Services section for further details on setting up a grid job.